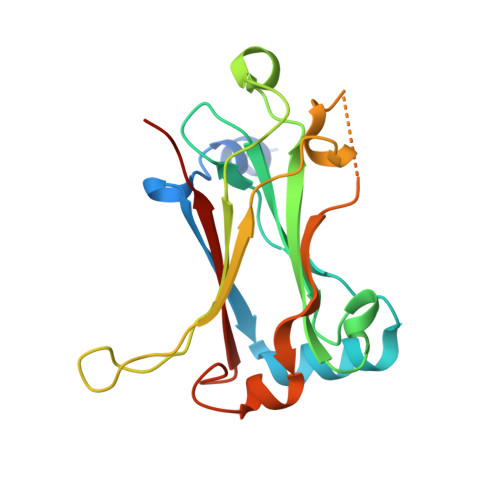
Structure of fish TRAF4 and its implication in TRAF4-mediated immune cell and platelet signaling.
Kim, C.M., Jang, H., Hong, E., Lee, J.H., Park, H.H.(2023) Fish Shellfish Immunol 132: 108462-108462
- PubMed: 36455779 Search on PubMed
- DOI: https://doi.org/10.1016/j.fsi.2022.108462
- Primary Citation Related Structures:
7YPC - PubMed Abstract:
Due to an increasing interest in immunity and signal transduction in teleost fish, important key signaling molecules associated with the immune response, including TRAF molecules, have been recently cloned and characterized. To better understand the role of TRAF4 in fish immune signaling and compare it with the human system, our study cloned the TRAF4 gene from the Antarctic yellowbelly rockcod Notothenia coriiceps (ncTRAF4) and purified the protein. Here, we report the first crystal structure of teleost fish TRAF4. Based on biochemical characterization, our findings elucidated the mechanisms through which signaling molecules gain cold adaptivity. Additionally, we identified a platelet receptor GPIbβ homolog in N. coriiceps (ncGPIbβ) and found that the "RRFERLFKEARRTS" region of this homolog directly binds to ncTRAF4, indicating that ncTRAF4 also recognizes the "RLXA" motif for receptor interactions and further TARF4-mediated cellular signaling. Collectively, our findings provide novel insights into the mechanisms of TRAF4-mediated immune cell and platelet signaling in fish and the structural flexibility-mediated cold adaptiveness of signaling molecules.
- College of Pharmacy, Chung-Ang University, Seoul, 06974, Republic of Korea.
Organizational Affiliation: