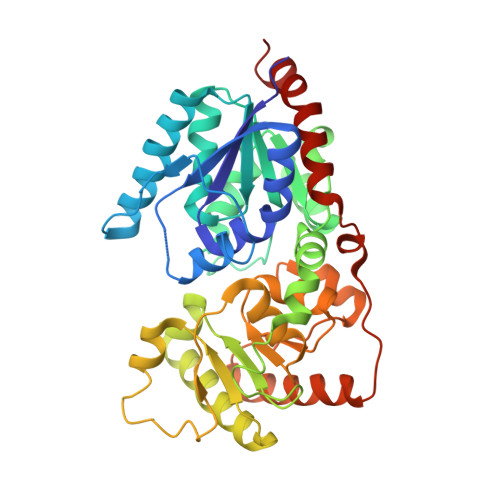
Crystal structure of the capsular polysaccharide-synthesis enzyme CapG from Staphylococcus aureus.
Tien, N., Ho, C.Y., Lai, S.J., Lin, Y.C., Yang, C.S., Wang, Y.C., Huang, W.C., Chen, Y., Chang, J.J.(2022) Acta Crystallogr F Struct Biol Commun 78: 378-385
- PubMed: 36322423 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X22008743
- Primary Citation Related Structures:
7YA2 - PubMed Abstract:
Bacterial capsular polysaccharides provide protection against environmental stress and immune evasion from the host immune system, and are therefore considered to be attractive therapeutic targets for the development of anti-infectious reagents. Here, we focused on CapG, one of the key enzymes in the synthesis pathway of capsular polysaccharides type 5 (CP5) from the opportunistic pathogen Staphylococcus aureus. SaCapG catalyses the 2-epimerization of UDP-N-acetyl-D-talosamine (UDP-TalNAc) to UDP-N-acetyl-D-fucosamine (UDP-FucNAc), which is one of the nucleotide-activated precursors for the synthesis of the trisaccharide repeating units of CP5. Here, the cloning, expression and purification of recombinant SaCapG are reported. After extensive efforts, single crystals of SaCapG were successfully obtained which belonged to space group C2 and exhibited unit-cell parameters a = 302.91, b = 84.34, c = 145.09 Å, β = 110.65°. The structure was solved by molecular replacement and was refined to 3.2 Å resolution. The asymmetric unit revealed a homohexameric assembly of SaCapG, which was consistent with gel-filtration analysis. Structural comparison with UDP-N-acetyl-D-glucosamine 2-epimerase from Methanocaldococcus jannaschii identified α2, the α2-α3 loop and α10 as a gate-regulated switch controlling substrate entry and/or product release.
- Department of Laboratory Medicine, China Medical University Hospital, Taichung, Taiwan.
Organizational Affiliation: