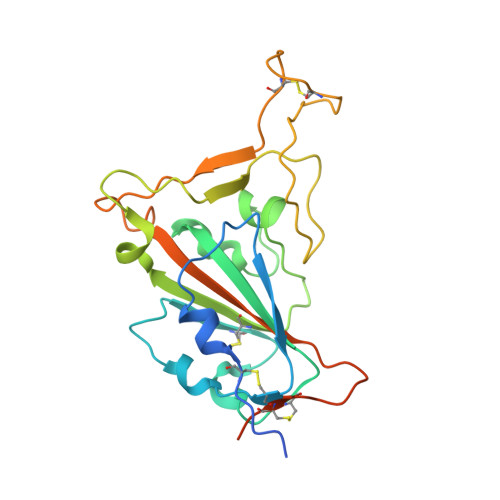
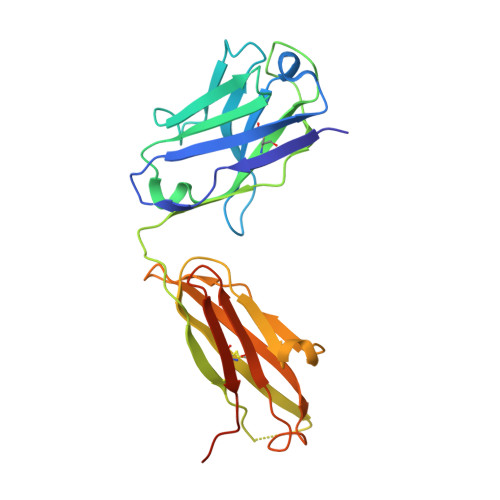
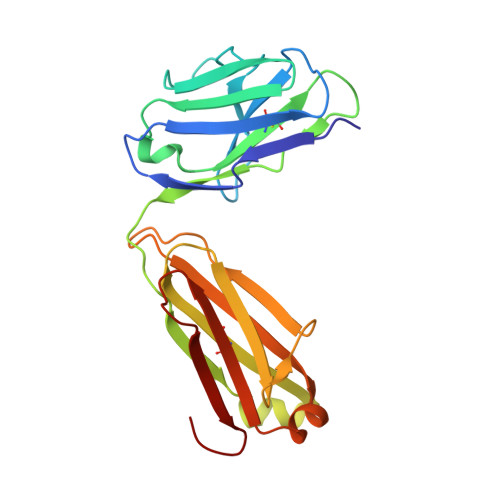
Defining a de novo non-RBM antibody as RBD-8 and its synergistic rescue of immune-evaded antibodies to neutralize Omicron SARS-CoV-2.
Rao, X., Zhao, R., Tong, Z., Guo, S., Peng, W., Liu, K., Li, S., Wu, L., Tong, J., Chai, Y., Han, P., Wang, F., Jia, P., Li, Z., Zhao, X., Li, D., Zhang, R., Zhang, X., Zou, W., Li, W., Wang, Q., Gao, G.F., Wu, Y., Dai, L., Gao, F.(2023) Proc Natl Acad Sci U S A 120: e2314193120-e2314193120
- PubMed: 38109549 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2314193120
- Primary Citation Related Structures:
7Y3N, 7Y3O, 8HRD - PubMed Abstract:
Currently, monoclonal antibodies (MAbs) targeting the SARS-CoV-2 receptor binding domain (RBD) of spike (S) protein are classified into seven classes based on their binding epitopes. However, most of these antibodies are seriously impaired by SARS-CoV-2 Omicron and its subvariants, especially the recent BQ.1.1, XBB and its derivatives. Identification of broadly neutralizing MAbs against currently circulating variants is imperative. In this study, we identified a "breathing" cryptic epitope in the S protein, named as RBD-8. Two human MAbs, BIOLS56 and IMCAS74, were isolated recognizing this epitope with broad neutralization abilities against tested sarbecoviruses, including SARS-CoV, pangolin-origin coronaviruses, and all the SARS-CoV-2 variants tested (Omicron BA.4/BA.5, BQ.1.1, and XBB subvariants). Searching through the literature, some more RBD-8 MAbs were defined. More importantly, BIOLS56 rescues the immune-evaded antibody, RBD-5 MAb IMCAS-L4.65, by making a bispecific MAb, to neutralize BQ.1 and BQ.1.1, thereby producing an MAb to cover all the currently circulating Omicron subvariants. Structural analysis reveals that the neutralization effect of RBD-8 antibodies depends on the extent of epitope exposure, which is affected by the angle of antibody binding and the number of up-RBDs induced by angiotensin-converting enzyme 2 binding. This cryptic epitope which recognizes non- receptor binding motif (non-RBM) provides guidance for the development of universal therapeutic antibodies and vaccines against COVID-19.
- Laboratory of Protein Engineering and Vaccines, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.
Organizational Affiliation: