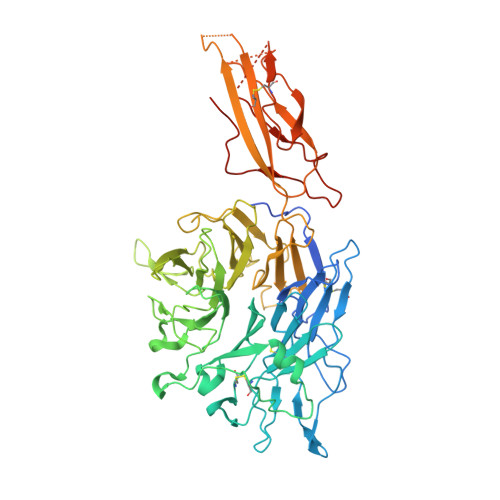
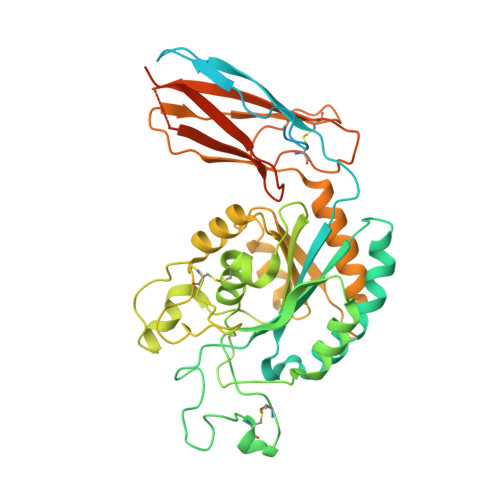
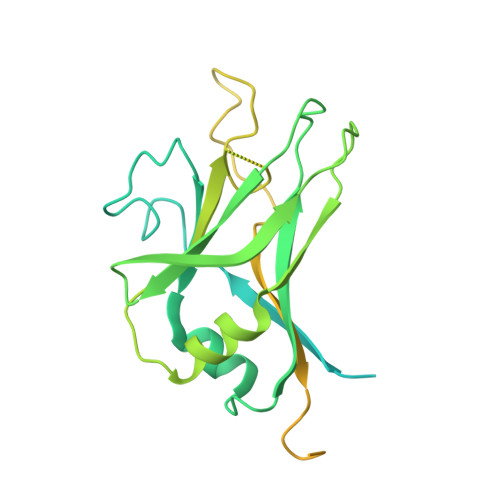
Specificity of TGF-beta 1 signal designated by LRRC33 and integrin alpha V beta 8.
Duan, Z., Lin, X., Wang, L., Zhen, Q., Jiang, Y., Chen, C., Yang, J., Lee, C.H., Qin, Y., Li, Y., Zhao, B., Wang, J., Zhang, Z.(2022) Nat Commun 13: 4988-4988
- PubMed: 36008481
- DOI: https://doi.org/10.1038/s41467-022-32655-9
- Primary Citation Related Structures:
7Y1R, 7Y1T - PubMed Abstract:
Myeloid lineage cells present the latent form of transforming growth factor-β1 (L-TGF-β1) to the membrane using an anchor protein LRRC33. Integrin α V β 8 activates extracellular L-TGF-β1 to trigger the downstream signaling functions. However, the mechanism designating the specificity of TGF-β1 presentation and activation remains incompletely understood. Here, we report cryo-EM structures of human L-TGF-β1/LRRC33 and integrin α V β 8 /L-TGF-β1 complexes. Combined with biochemical and cell-based analyses, we demonstrate that LRRC33 only presents L-TGF-β1 but not the -β2 or -β3 isoforms due to difference of key residues on the growth factor domains. Moreover, we reveal a 2:2 binding mode of integrin α V β 8 and L-TGF-β1, which shows higher avidity and more efficient L-TGF-β1 activation than previously reported 1:2 binding mode. We also uncover that the disulfide-linked loop of the integrin subunit β 8 determines its exquisite affinity to L-TGF-β1. Together, our findings provide important insights into the specificity of TGF-β1 signaling achieved by LRRC33 and integrin α V β 8 .
- State Key Laboratory of Membrane Biology, Center for Life Sciences, School of Life Sciences, Peking University, 100871, Beijing, China.
Organizational Affiliation: