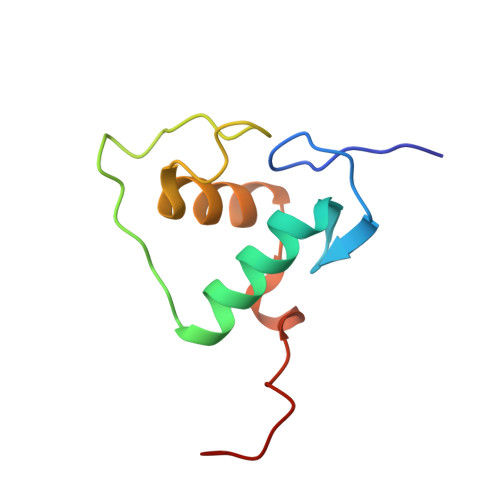
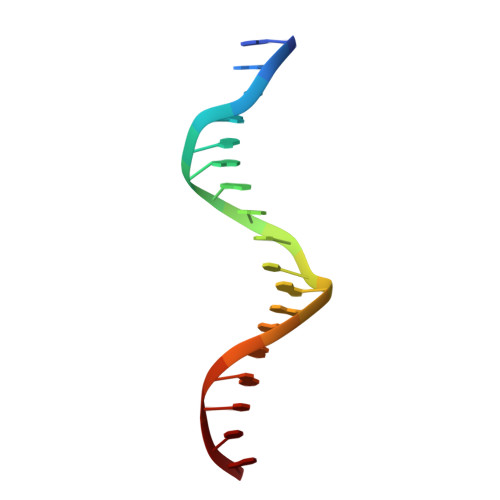
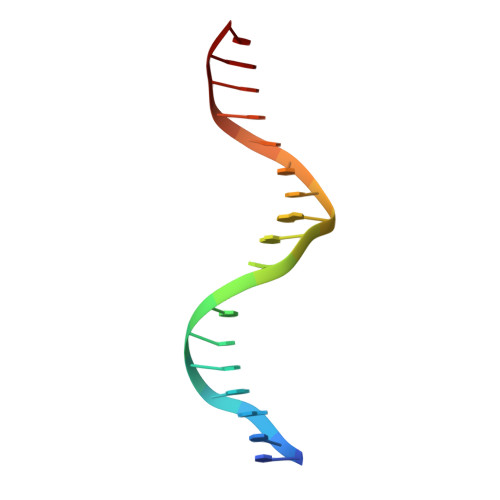
Structures of human TR4LBD-JAZF1 and TR4DBD-DNA complexes reveal the molecular basis of transcriptional regulation.
Liu, Y., Ma, L., Li, M., Tian, Z., Yang, M., Wu, X., Wang, X., Shang, G., Xie, M., Chen, Y., Liu, X., Jiang, L., Wu, W., Xu, C., Xia, L., Li, G., Dai, S., Chen, Z.(2023) Nucleic Acids Res 51: 1443-1457
- PubMed: 36651297 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac1259
- Primary Citation Related Structures:
7XV6, 7XV8, 7XV9, 7XVA - PubMed Abstract:
Testicular nuclear receptor 4 (TR4) modulates the transcriptional activation of genes and plays important roles in many diseases. The regulation of TR4 on target genes involves direct interactions with DNA molecules via the DNA-binding domain (DBD) and recruitment of coregulators by the ligand-binding domain (LBD). However, their regulatory mechanisms are unclear. Here, we report high-resolution crystal structures of TR4DBD, TR4DBD-DNA complexes and the TR4LBD-JAZF1 complex. For DNA recognition, multiple factors come into play, and a specific mutual selectivity between TR4 and target genes is found. The coactivators SRC-1 and CREBBP can bind at the interface of TR4 originally occupied by the TR4 activation function region 2 (AF-2); however, JAZF1 suppresses the binding through a novel mechanism. JAZF1 binds to an unidentified surface of TR4 and stabilizes an α13 helix never reported in the nuclear receptor family. Moreover, the cancer-associated mutations affect the interactions and the transcriptional activation of TR4 in vitro and in vivo, respectively. Overall, our results highlight the crucial role of DNA recognition and a novel mechanism of how JAZF1 reinforces the autorepressed conformation and influences the transcriptional activation of TR4, laying out important structural bases for drug design for a variety of diseases, including diabetes and cancers.
- State Key Laboratory of Agrobiotechnology and Beijing Advanced Innovation Center for Food Nutrition and Human Health, College of Biological Sciences, China Agricultural University, Beijing 100193, China.
Organizational Affiliation: