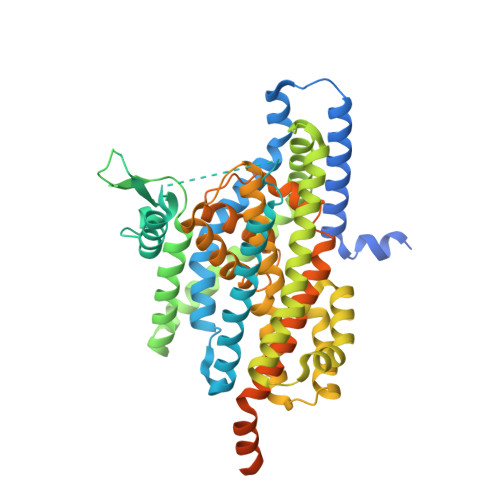
Structural basis of ligand binding modes of human EAAT2.
Zhang, Z., Chen, H., Geng, Z., Yu, Z., Li, H., Dong, Y., Zhang, H., Huang, Z., Jiang, J., Zhao, Y.(2022) Nat Commun 13: 3329-3329
- PubMed: 35680945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-31031-x
- Primary Citation Related Structures:
7XR4, 7XR6 - PubMed Abstract:
In the central nervous system (CNS), excitatory amino acid transporters (EAATs) mediate the uptake of excitatory neurotransmitter glutamate and maintain its low concentrations in the synaptic cleft for avoiding neuronal cytotoxicity. Dysfunction of EAATs can lead to many psychiatric diseases. Here we report cryo-EM structures of human EAAT2 in an inward-facing conformation, in the presence of substrate glutamate or selective inhibitor WAY-213613. The glutamate is coordinated by extensive hydrogen bonds and further stabilized by HP2. The inhibitor WAY-213613 occupies a similar binding pocket to that of the substrate glutamate. Upon association with the WAY-213613, the HP2 undergoes a substantial conformational change, and in turn stabilizes the inhibitor binding by forming hydrophobic interactions. Electrophysiological experiments elucidate that the unique S441 plays pivotal roles in the binding of hEAAT2 with glutamate or WAY-213613, and the I464-L467-V468 cluster acts as a key structural determinant for the selective inhibition of this transporter by WAY-213613.
- Department of Microbiology and Biotechnology, College of Life Sciences, Northeast Agricultural University, No. 600 Changjiang Road, Xiangfang District, Harbin, 150030, China.
Organizational Affiliation: