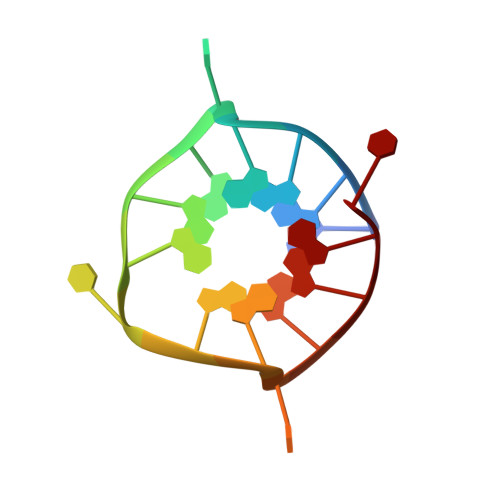
Crystal structures of an HIV-1 integrase aptamer: Formation of a water-mediated A.G.G.G.G pentad in an interlocked G-quadruplex.
Ngo, K.H., Liew, C.W., Lattmann, S., Winnerdy, F.R., Phan, A.T.(2022) Biochem Biophys Res Commun 613: 153-158
- PubMed: 35561583 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2022.04.020
- Primary Citation Related Structures:
7X7G, 7XDH, 7XH9, 7XHD, 7XIE - PubMed Abstract:
93del is a 16-nucleotide G-quadruplex-forming aptamer which can inhibit the activity of the HIV-1 integrase enzyme at nanomolar concentration. Previous structural analyses of 93del using NMR spectroscopy have shown that the aptamer forms an interlocked G-quadruplex structure in K + solution. Due to its exceptional stability and unique topology, 93del has been used in many different studies involving DNA G-quadruplexes, such as DNA aptamer and multimer design, as well as DNA fluorescence research. To gain further insights on the structure of this unique aptamer, we have determined several high-resolution crystal structures of 93del and its variants. While confirming the overall dimeric interlocked G-quadruplex folding topology previously determined by NMR, our results reveal important detailed structural information, particularly the formation of a water-mediated A•G•G•G•G pentad. These insights allow us to better understand the formation of various structural elements in G-quadruplexes and should be useful for designing and manipulating G-quadruplex scaffolds with desired properties.
- School of Physical and Mathematical Sciences, Nanyang Technological University, Singapore, 637371, Singapore.
Organizational Affiliation: