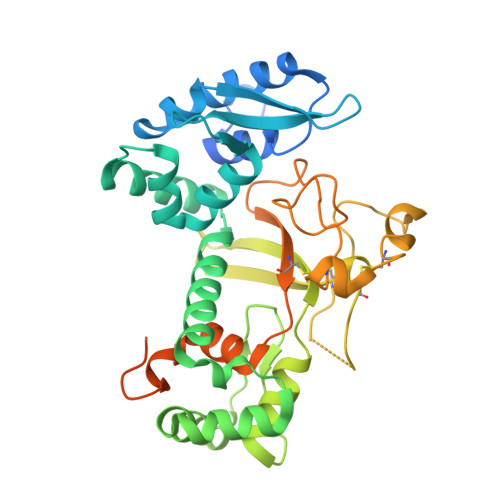
Structural Characterization of Per Os Infectivity Factor 5 (PIF5) Reveals the Essential Role of Intramolecular Interactions in Baculoviral Oral Infectivity.
Li, Z., Zhang, H., Li, Z., Fu, Y., Wang, X., Li, J., Wang, K., Wang, Z., Zhang, T., Wang, M., Hu, Z., Cao, S.(2022) J Virol 96: e0080622-e0080622
- PubMed: 35862697 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/jvi.00806-22
- Primary Citation Related Structures:
7X77 - PubMed Abstract:
Baculoviruses initiate oral infection in the highly alkaline midgut of insects via a group of envelope proteins called per os infectivity factors (PIFs). To date, no high-resolution structural information has been reported for any PIF. Here, we present the crystal structure of the PIF5 ectodomain (PIF5e) from Autographa californica multiple nucleopolyhedrovirus (AcMNPV) at a 2.2-Å resolution. It revealed an open cavity between the N-terminal E1 domain and the C-terminal E2 domain and a cysteine-rich region with three pairs of disulfide bonds in the E2 domain. Multiple conserved intramolecular interactions within PIF5 are essential for maintaining its tertiary structure. Two conserved arginines (Arg8 and Arg74) play critical roles in E1-E2 interactions, and mutagenesis analysis supported their crucial role in oral infection. Importantly, the reduction in the oral infectivity of the Arg8, Arg74, or cysteine mutant viruses was related to the proteolytic cleavage of PIF5 by the endogenous protease embedded in occlusion bodies during alkaline treatment. This suggested that the structural stability of PIF5 under physiological conditions in the insect midgut is critical for baculoviral oral infectivity. IMPORTANCE Per os infection mediated by PIFs is the highly complex mechanism by which baculoviruses initiate infection in insects. Previous studies revealed that multiple PIF proteins form a large PIF complex on the envelope of virions, while PIF5 functions independently of the PIF complex. Here, we report the crystal structure of AcMNPV PIF5e, which, to our knowledge, is the first atomic structure reported for a PIF protein. The structure revealed the precise locations of three previously proposed disulfide bonds and other conserved intramolecular interactions, which are important for the structural stability of PIF5 and are also essential for oral infectivity. These findings advance our understanding of the molecular mechanism of baculovirus oral infection under alkaline conditions.
- CAS Key Laboratory of Special Pathogens, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, Wuhan, People's Republic of China.
Organizational Affiliation: