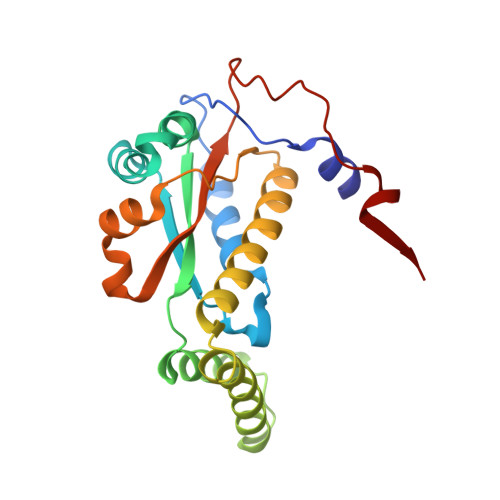
Structural basis for the transformation of the traditional medicine berberine by bacterial nitroreductase.
Wen, H.Y., Pan, L.B., Ma, S.R., Yang, X.Y., Hu, J.C., Zhao, H.F., Gao, Z.Q., Dong, Y.H., Wang, Y., Zhang, H.(2022) Acta Crystallogr D Struct Biol 78: 1273-1282
- PubMed: 36189746 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798322008373
- Primary Citation Related Structures:
7X32 - PubMed Abstract:
The bacterial nitroreductases (NRs) NfsB and NfsA are conserved homodimeric FMN-dependent flavoproteins that are responsible for the reduction of nitroaromatic substrates. Berberine (BBR) is a plant-derived isoquinoline alkaloid with a large conjugated ring system that is widely used in the treatment of various diseases. It was recently found that the gut microbiota convert BBR into dihydroberberine (dhBBR, the absorbable form) mediated by bacterial NRs. The molecular basis for the transformation of BBR by the gut microbiota remains unclear. Here, kinetic studies showed that NfsB from Escherichia coli (EcNfsB), rather than EcNfsA, is responsible for the conversion of BBR to dhBBR in spite of a low reaction rate. The crystal structure of the EcNfsB-BBR complex showed that BBR binds into the active pocket at the dimer interface, and its large conjugated plane stacks above the plane of the FMN cofactor in a nearly parallel orientation. BBR is mainly stabilized by π-stacking interactions with both neighboring aromatic residues and FMN. Structure-based mutagenesis studies further revealed that the highly conserved Phe70 and Phe199 are important residues for the conversion of BBR. The structure revealed that the C6 atom of BBR (which receives the hydride) is ∼7.5 Å from the N5 atom of FMN (which donates the hydride), which is too distant for hydride transfer. Notably, several well ordered water molecules make hydrogen-bond/van der Waals contacts with the N1 atom of BBR in the active site, which probably donate protons in conjunction with electron transfer from FMN. The structure-function studies revealed the mechanism for the recognition and binding of BBR by bacterial NRs and may help to understand the conversion of BBR by the gut microbiota.
- School of Life Sciences, University of Science and Technology of China, Hefei 230027, People's Republic of China.
Organizational Affiliation: