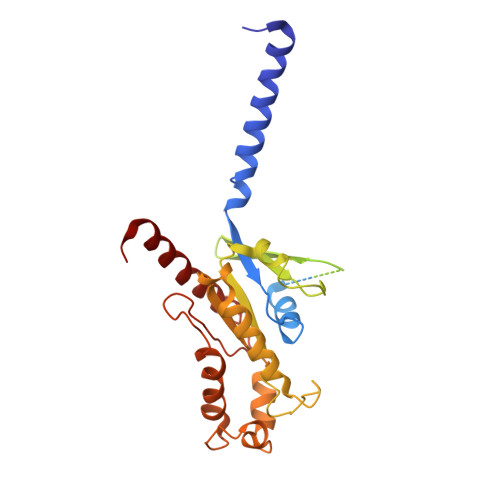
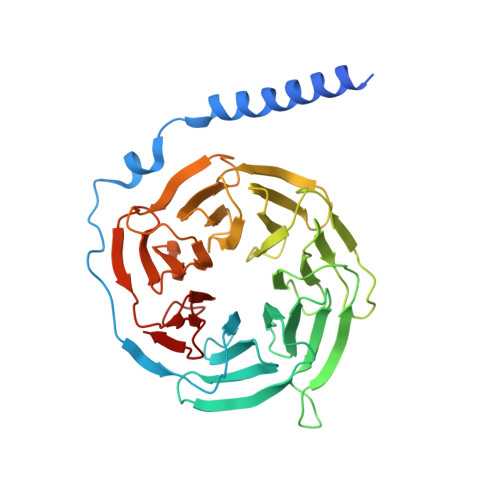
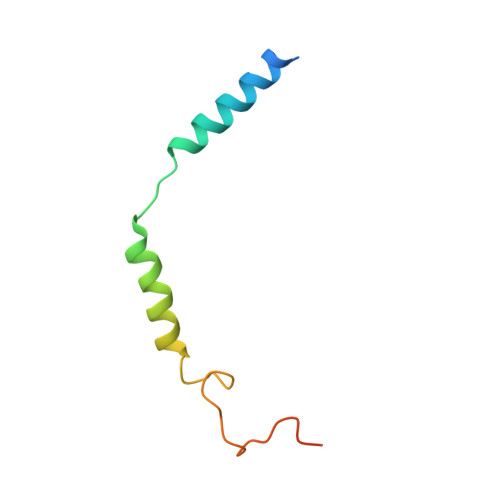
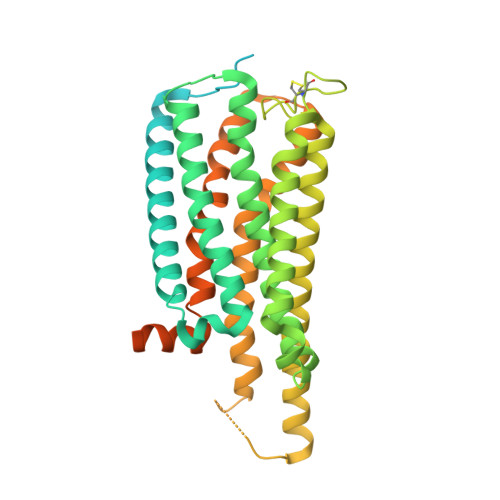
Molecular mechanism of allosteric modulation for the cannabinoid receptor CB1.
Yang, X., Wang, X., Xu, Z., Wu, C., Zhou, Y., Wang, Y., Lin, G., Li, K., Wu, M., Xia, A., Liu, J., Cheng, L., Zou, J., Yan, W., Shao, Z., Yang, S.(2022) Nat Chem Biol 18: 831-840
- PubMed: 35637350 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-022-01038-y
- Primary Citation Related Structures:
7FEE, 7WV9 - PubMed Abstract:
Given the promising clinical value of allosteric modulators of G protein-coupled-receptors (GPCRs), mechanistic understanding of how these modulators alter GPCR function is of significance. Here, we report the crystallographic and cryo-electron microscopy structures of the cannabinoid receptor CB1 bound to the positive allosteric modulator (PAM) ZCZ011. These structures show that ZCZ011 binds to an extrahelical site in the transmembrane 2 (TM2)-TM3-TM4 surface. Through (un)biased molecular dynamics simulations and mutagenesis experiments, we show that TM2 rearrangement is critical for the propagation of allosteric signals. ZCZ011 exerts a PAM effect by promoting TM2 rearrangement in favor of receptor activation and increasing the population of receptors that adopt an active conformation. In contrast, ORG27569, a negative allosteric modulator (NAM) of CB1, also binds to the TM2-TM3-TM4 surface and exerts a NAM effect by impeding the TM2 rearrangement. Our findings fill a gap in the understanding of CB1 allosteric regulation and could guide the rational design of CB1 allosteric modulators.
- State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, Chengdu, Sichuan, China.
Organizational Affiliation: