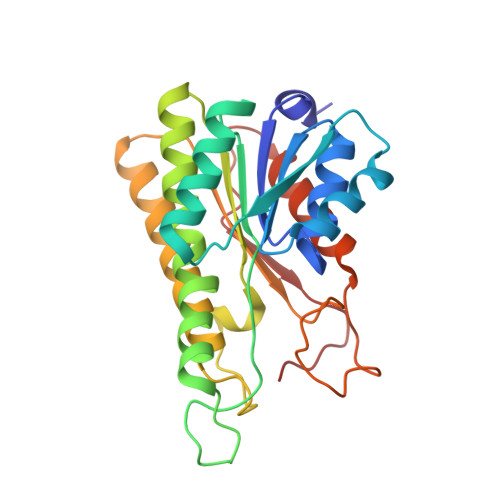
Crystal structure and molecular characterization of NADP + -farnesol dehydrogenase from cotton bollworm, Helicoverpaarmigera.
Kumar, R., Das, J., Mahto, J.K., Sharma, M., Vivek, S., Kumar, P., Sharma, A.K.(2022) Insect Biochem Mol Biol 147: 103812-103812
- PubMed: 35820537 Search on PubMed
- DOI: https://doi.org/10.1016/j.ibmb.2022.103812
- Primary Citation Related Structures:
7W61 - PubMed Abstract:
Farnesol dehydrogenase (FDL) orchestrates the oxidation reaction catalyzing farnesol to farnesal, a key step in the juvenile hormone (JH) biosynthesis pathway of insects and hence, represents a lucrative target for developing insect growth regulators (IGRs). However, information on the structural and functional characterization of JH-specific farnesol dehydrogenase in insects remains elusive. Herein, we identified a transcript that encodes farnesol dehydrogenase (HaFDL) from Helicoverpa armigera, a major pest of cotton. The investigations of molecular assembly, biochemical analysis and spatio-temporal expression profiling showed that HaFDL exists as a soluble homo-tetrameric form, exhibits a broad substrate affinity and is involved in the JH-specific farnesol oxidation in H. armigera. Additionally, the study presents the first crystal structure of the HaFDL-NADP enzyme complex determined at 1.6 Å resolution. Structural analysis revealed that HaFDL belongs to the NADP-specific cP2 subfamily of the classical short-chain dehydrogenase/reductase (SDR) family and exhibits typical structural features of those enzymes including the conserved nucleotide-binding Rossman-fold. The isothermal titration calorimetry (ITC) showed a high binding affinity (dissociation constant, Kd, 3.43 μM) of NADP to the enzyme. Comparative structural analysis showed a distinct substrate-binding pocket (SBP) loop with a spacious and hydrophobic substrate-binding pocket in HaFDL, consistent with the biochemically observed promiscuous substrate specificity. Finally, based on the crystal structure, substrate modeling and structural comparison with homologs, a two-step reaction mechanism is proposed. Overall, the findings significantly impact and contribute to our understanding of farnesol dehydrogenase functional properties in JH biosynthesis in H. armigera.
- Department of Biosciences and Bioengineering, Indian Institute of Technology Roorkee, Roorkee, 247 667, India; ICAR-Central Institute for Cotton Research, Nagpur, India.
Organizational Affiliation: