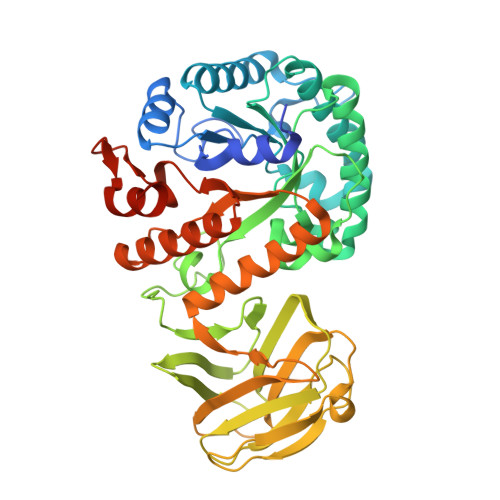
Synergic action of an inserted carbohydrate-binding module in a glycoside hydrolase family 5 endoglucanase.
Ye, T.J., Huang, K.F., Ko, T.P., Wu, S.H.(2022) Acta Crystallogr D Struct Biol 78: 633-646
- PubMed: 35503211
- DOI: https://doi.org/10.1107/S2059798322002601
- Primary Citation Related Structures:
7VT4, 7VT5, 7VT6, 7VT7, 7VT8 - PubMed Abstract:
Most known cellulase-associated carbohydrate-binding modules (CBMs) are attached to the N- or C-terminus of the enzyme or are expressed separately and assembled into multi-enzyme complexes (for example to form cellulosomes), rather than being an insertion into the catalytic domain. Here, by solving the crystal structure, it is shown that MtGlu5 from Meiothermus taiwanensis WR-220, a GH5-family endo-β-1,4-glucanase (EC 3.2.1.4), has a bipartite architecture consisting of a Cel5A-like catalytic domain with a (β/α) 8 TIM-barrel fold and an inserted CBM29-like noncatalytic domain with a β-jelly-roll fold. Deletion of the CBM significantly reduced the catalytic efficiency of MtGlu5, as determined by isothermal titration calorimetry using inactive mutants of full-length and CBM-deleted MtGlu5 proteins. Conversely, insertion of the CBM from MtGlu5 into TmCel5A from Thermotoga maritima greatly enhanced the substrate affinity of TmCel5A. Bound sugars observed between two tryptophan side chains in the catalytic domains of active full-length and CBM-deleted MtGlu5 suggest an important stacking force. The synergistic action of the catalytic domain and CBM of MtGlu5 in binding to single-chain polysaccharides was visualized by substrate modeling, in which additional surface tryptophan residues were identified in a cross-domain groove. Subsequent site-specific mutagenesis results confirmed the pivotal role of several other tryptophan residues from both domains of MtGlu5 in substrate binding. These findings reveal a way to incorporate a CBM into the catalytic domain of an existing enzyme to make a robust cellulase.
- Institute of Biological Chemistry, Academia Sinica, Taipei 115, Taiwan.
Organizational Affiliation: