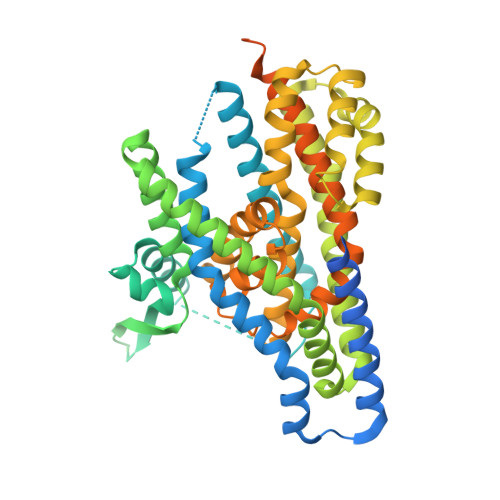
Structural insights into inhibitory mechanism of human excitatory amino acid transporter EAAT2.
Kato, T., Kusakizako, T., Jin, C., Zhou, X., Ohgaki, R., Quan, L., Xu, M., Okuda, S., Kobayashi, K., Yamashita, K., Nishizawa, T., Kanai, Y., Nureki, O.(2022) Nat Commun 13: 4714-4714
- PubMed: 35953475
- DOI: https://doi.org/10.1038/s41467-022-32442-6
- Primary Citation of Related Structures:
7VR7, 7VR8 - PubMed Abstract:
Glutamate is a pivotal excitatory neurotransmitter in mammalian brains, but excessive glutamate causes numerous neural disorders. Almost all extracellular glutamate is retrieved by the glial transporter, Excitatory Amino Acid Transporter 2 (EAAT2), belonging to the SLC1A family. However, in some cancers, EAAT2 expression is enhanced and causes resistance to therapies by metabolic disturbance. Despite its crucial roles, the detailed structural information about EAAT2 has not been available. Here, we report cryo-EM structures of human EAAT2 in substrate-free and selective inhibitor WAY213613-bound states at 3.2 Å and 2.8 Å, respectively. EAAT2 forms a trimer, with each protomer consisting of transport and scaffold domains. Along with a glutamate-binding site, the transport domain possesses a cavity that could be disrupted during the transport cycle. WAY213613 occupies both the glutamate-binding site and cavity of EAAT2 to interfere with its alternating access, where the sensitivity is defined by the inner environment of the cavity. We provide the characterization of the molecular features of EAAT2 and its selective inhibition mechanism that may facilitate structure-based drug design for EAAT2.
- Department of Biological Science, Graduate School of Science, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: