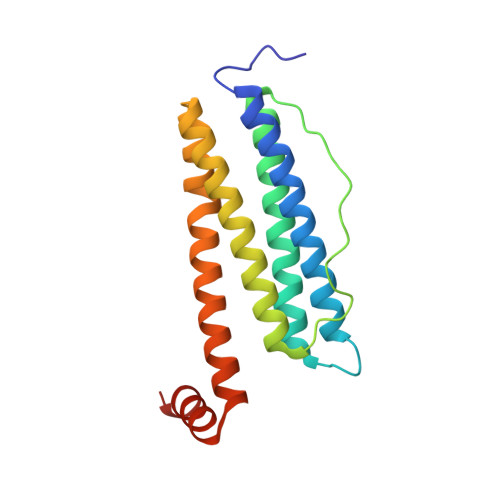
Design of a gold clustering site in an engineered apo-ferritin cage.
Lu, C., Maity, B., Peng, X., Ito, N., Abe, S., Sheng, X., Ueno, T., Lu, D.(2022) Commun Chem 5: 39-39
- PubMed: 36697940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42004-022-00651-1
- Primary Citation Related Structures:
7VIO, 7VIP, 7VIQ, 7VIR, 7VIS, 7VIT, 7VIU - PubMed Abstract:
Water-soluble and biocompatible protein-protected gold nanoclusters (Au NCs) hold great promise for numerous applications. However, design and precise regulation of their structure at an atomic level remain challenging. Herein, we have engineered and constructed a gold clustering site at the 4-fold symmetric axis channel of the apo-ferritin cage. Using a series of X-ray crystal structures, we evaluated the stepwise accumulation process of Au ions into the cage and the formation of a multinuclear Au cluster in our designed cavity. We also disclosed the role of key residues in the metal accumulation process. X-ray crystal structures in combination with quantum chemical (QC) calculation revealed a unique Au clustering site with up to 12 Au atoms positions in the cavity. Moreover, the structure of the gold nanocluster was precisely tuned by the dosage of the Au precursor. As the gold concentration increases, the number of Au atoms position at the clustering site increases from 8 to 12, and a structural rearrangement was observed at a higher Au concentration. Furthermore, the binding affinity order of the four Au binding sites on apo-ferritin was unveiled with a stepwise increase of Au precursor concentration.
- Department of Chemical Engineering, Tsinghua University, Beijing, 100-084, China.
Organizational Affiliation: