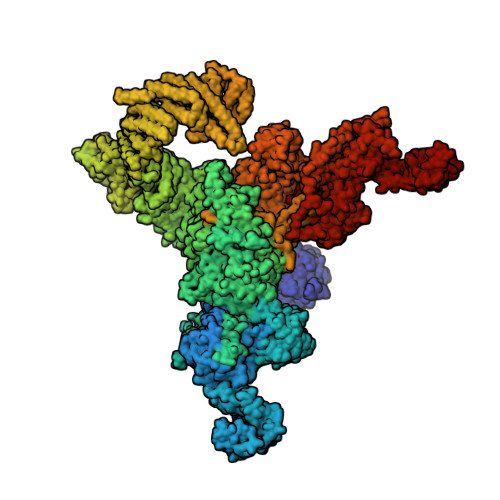
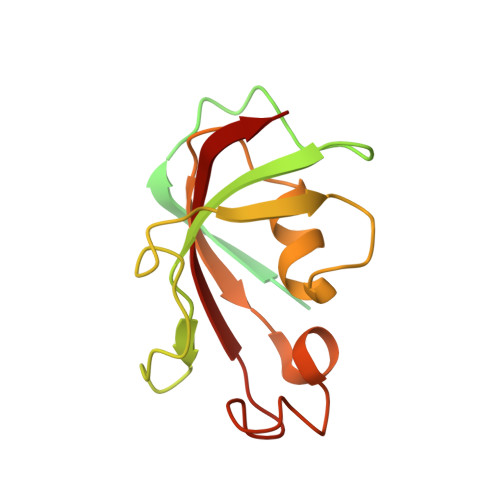
Molecular basis for gating of cardiac ryanodine receptor explains the mechanisms for gain- and loss-of function mutations
Kobayashi, T., Tsutsumi, A., Kurebayashi, N., Saito, K., Kodama, M., Sakurai, T., Kikkawa, M., Murayama, T., Ogawa, H.(2022) Nat Commun 13: 2821