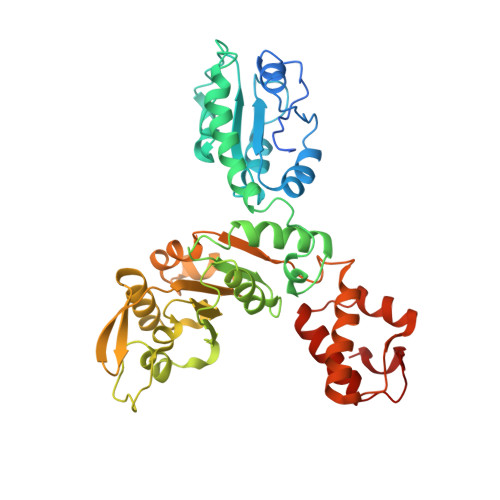
Crystal Structure of VpsR Revealed Novel Dimeric Architecture and c-di-GMP Binding Site: Mechanistic Implications in Oligomerization, ATPase Activity and DNA Binding.
Chakrabortty, T., Roy Chowdhury, S., Ghosh, B., Sen, U.(2022) J Mol Biology 434: 167354-167354
- PubMed: 34774564 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2021.167354
- Primary Citation Related Structures:
7V2B, 7V2V, 7V3W, 7V4E - PubMed Abstract:
VpsR, the master regulator of biofilm formation in Vibrio cholerae, is an atypical NtrC1 type bEBP lacking residues essential for σ 54 -RNAP binding and REC domain phosphorylation. Moreover, transcription from P vpsL , a promoter of biofilm biosynthesis, has been documented in presence of σ 70 -RNAP/VpsR/c-di-GMP complex. It was proposed that c-di-GMP and VpsR together form an active transcription complex with σ 70 -RNAP. However, the impact of c-di-GMP imparted on VpsR that leads to transcription activation with σ 70 -RNAP remained elusive, largely due to the lack of the structure of VpsR and knowledge about c-di-GMP:VpsR interactions. In this direction we have solved the crystal structure of VpsR RA , containing REC and AAA + domains, in apo, AMPPNP/GMPPNP and c-di-GMP bound states. Structures of VpsR RA unveiled distinctive REC domain orientation that leads to a novel dimeric association and noncanonical ATP/GTP binding. Moreover, we have demonstrated that at physiological pH VpsR remains as monomer having no ATPase activity but c-di-GMP imparted cooperativity to convert it to dimer with potent activity. Crystal structure of c-di-GMP:VpsR RA complex reveals that c-di-GMP binds near the C-terminal end of AAA + domain. Trp quenching studies on VpsR R , VpsR A , VpsR RA , VpsR AD with c-di-GMP additionally demonstrated that c-di-GMP could potentially bind VpsR D . We propose that c-di-GMP mediated tethering of VpsR D with VpsR A could likely favor generating the specific protein-DNA architecture for transcription activation.
- Crystallography and Molecular Biology Division, Saha Institute of Nuclear Physics, HBNI, 1/AF Bidhan Nagar, Kolkata 700064, India. Electronic address: https://twitter.com/@TulikaC02382598.
Organizational Affiliation: