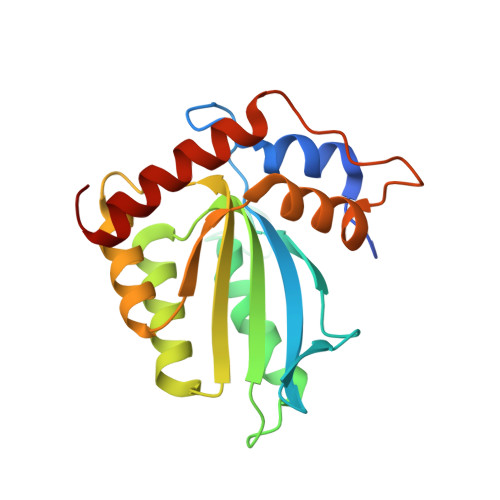
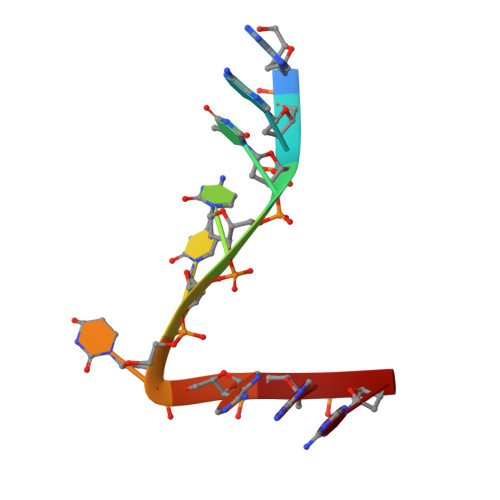
Structure of the catalytically active APOBEC3G bound to a DNA oligonucleotide inhibitor reveals tetrahedral geometry of the transition state.
Maiti, A., Hedger, A.K., Myint, W., Balachandran, V., Watts, J.K., Schiffer, C.A., Matsuo, H.(2022) Nat Commun 13: 7117-7117
- PubMed: 36402773 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34752-1
- Primary Citation Related Structures:
7UXD - PubMed Abstract:
APOBEC3 proteins (A3s) are enzymes that catalyze the deamination of cytidine to uridine in single-stranded DNA (ssDNA) substrates, thus playing a key role in innate antiviral immunity. However, the APOBEC3 family has also been linked to many mutational signatures in cancer cells, which has led to an intense interest to develop inhibitors of A3's catalytic activity as therapeutics as well as tools to study A3's biochemistry, structure, and cellular function. Recent studies have shown that ssDNA containing 2'-deoxy-zebularine (dZ-ssDNA) is an inhibitor of A3s such as A3A, A3B, and A3G, although the atomic determinants of this activity have remained unknown. To fill this knowledge gap, we determined a 1.5 Å resolution structure of a dZ-ssDNA inhibitor bound to active A3G. The crystal structure revealed that the activated dZ-H 2 O mimics the transition state by coordinating the active site Zn 2+ and engaging in additional stabilizing interactions, such as the one with the catalytic residue E259. Therefore, this structure allowed us to capture a snapshot of the A3's transition state and suggests that developing transition-state mimicking inhibitors may provide a new opportunity to design more targeted molecules for A3s in the future.
- Cancer Innovation Laboratory, Frederick National Laboratory for Cancer Research, Frederick, MD, USA.
Organizational Affiliation: