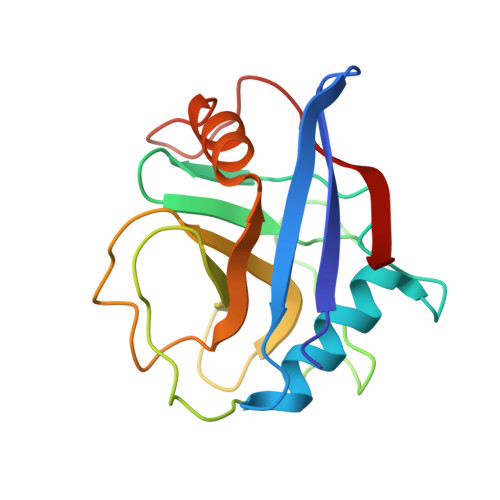
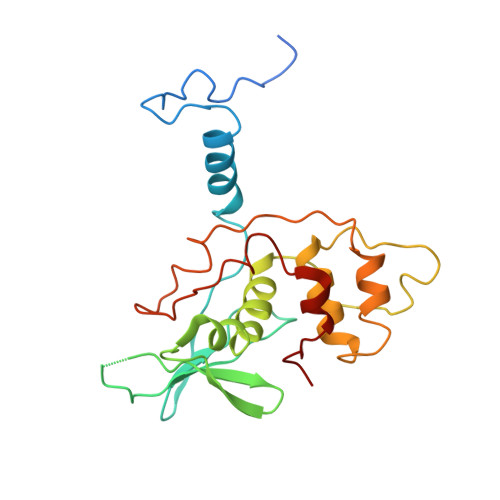
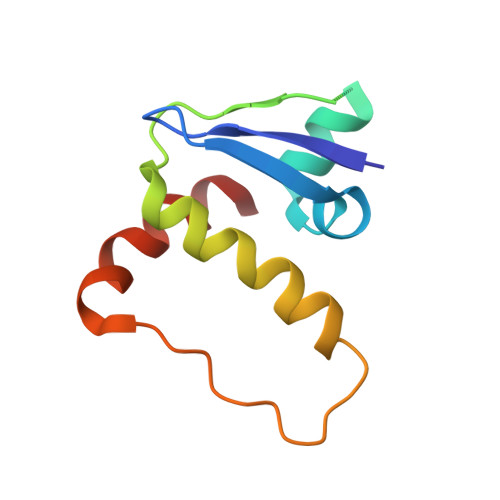
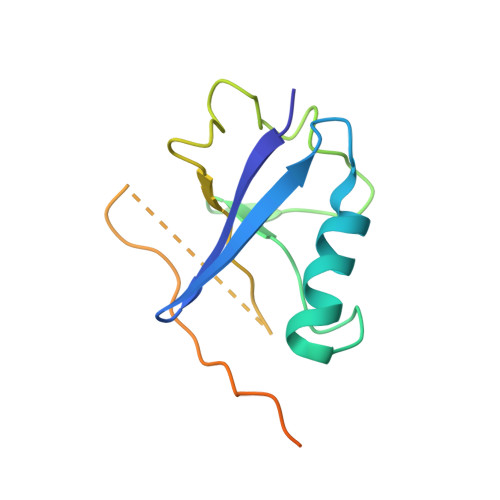
Structural basis for recruitment of host CypA and E3 ubiquitin ligase by maedi-visna virus Vif.
Hu, Y., Gudnadottir, R.B., Knecht, K.M., Arizaga, F., Jonsson, S.R., Xiong, Y.(2023) Sci Adv 9: eadd3422-eadd3422
- PubMed: 36638173 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.add3422
- Primary Citation Related Structures:
7UPN - PubMed Abstract:
Lentiviral Vif molecules target the host antiviral APOBEC3 proteins for destruction in cellular ubiquitin-proteasome pathways. Different lentiviral Vifs have evolved to use the same canonical E3 ubiquitin ligase complexes, along with distinct noncanonical host cofactors for their activities. Unlike primate lentiviral Vif, which recruits CBFβ as the noncanonical cofactor, nonprimate lentiviral Vif proteins have developed different cofactor recruitment mechanisms. Maedi-visna virus (MVV) sequesters CypA as the noncanonical cofactor for the Vif-mediated ubiquitination of ovine APOBEC3s. Here, we report the cryo-electron microscopy structure of MVV Vif in complex with CypA and E3 ligase components. The structure, along with our biochemical and functional analysis, reveals both conserved and unique structural elements of MVV Vif and its common and distinct interaction modes with various cognate cellular proteins, providing a further understanding of the evolutionary relationship between lentiviral Vifs and the molecular mechanisms by which they capture different host cofactors for immune evasion activities.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT, USA.
Organizational Affiliation: