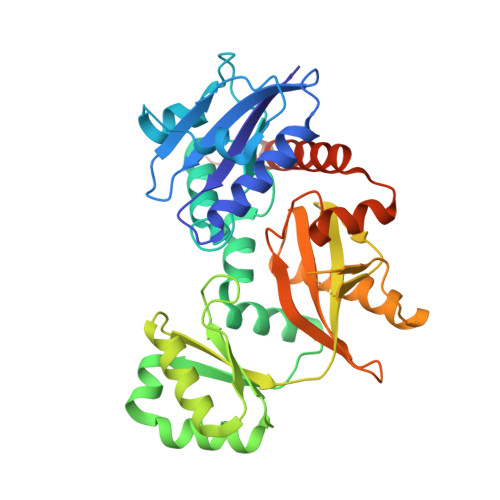
Structure-guided design and synthesis of ATP-competitive N-acyl-substituted sulfamide d-alanine-d-alanine ligase inhibitors.
Becker, R., Pederick, J.L., Dawes, E.G., Bruning, J.B., Abell, A.D.(2023) Bioorg Med Chem 96: 117509-117509
- PubMed: 37948922 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2023.117509
- Primary Citation Related Structures:
7U9K - PubMed Abstract:
d-Alanine-d-alanine ligase (Ddl) catalyses the ATP-dependent formation of d-Ala-d-Ala, a critical component in bacterial cell wall biosynthesis and is a validated target for new antimicrobial agents. Here, we describe the structure-guided design, synthesis, and evaluation of ATP-competitive N-acyl-substituted sulfamides 27-36, 42, 46, 47 as inhibitors of Staphylococcus aureus Ddl (SaDdl). A crystal structure of SaDdl complexed with ATP and d-Ala-d-Ala (PDB: 7U9K) identified ATP-mimetic 8 as an initial scaffold for further inhibitor design. Evaluation of 8 in SaDdl enzyme inhibition assays revealed the ability to reduce enzyme activity to 72 ± 8 % (IC 50 = 1.6 mM). The sulfamide linker of 8 was extended with 2-(4-methoxyphenyl)ethanol to give 29, to investigate further interactions with the d-Ala pocket of SaDdl, as predicted by molecular docking. This compound reduced enzyme activity to 89 ± 1 %, with replacement of the 4-methoxyphenyl group in 29 with alternative phenyl substituents (27, 28, 31-33, 35, 36) failing to significantly improve on this (80-89 % remaining enzyme activity). Exchanging these phenyl substituents with selected heterocycles (42, 46, 47) did improve activity, with the most active compound (42) reducing SaDdl activity to 70 ± 1 % (IC 50 = 1.7 mM), which compares favourably to the FDA-approved inhibitor d-cycloserine (DCS) (IC 50 = 0.1 mM). To the best of our knowledge, this is the first reported study of bisubstrate SaDdl inhibitors.
- Department of Chemistry, School of Physics, Chemistry and Earth Sciences, University of Adelaide, Adelaide, South Australia 5005, Australia; Institute for Photonics and Advanced Sensing, (IPAS), School of Biological Sciences, The University of Adelaide, Adelaide, South Australia 5005, Australia; Centre for Nanoscale BioPhotonics (CNBP), University of Adelaide, Adelaide, South Australia 5005, Australia.
Organizational Affiliation: