Redesign of Rifamycin Antibiotics to Overcome ADP-Ribosylation-Mediated Resistance.
Lan, T., Ganapathy, U.S., Sharma, S., Ahn, Y.M., Zimmerman, M., Molodtsov, V., Hegde, P., Gengenbacher, M., Ebright, R.H., Dartois, V., Freundlich, J.S., Dick, T., Aldrich, C.C.(2022) Angew Chem Int Ed Engl 61: e202211498-e202211498
- PubMed: 36222275 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202211498
- Primary Citation Related Structures:
7U22 - PubMed Abstract:
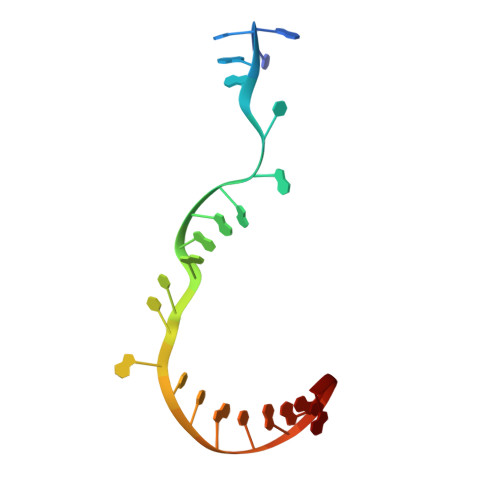
Rifamycin antibiotics are a valuable class of antimicrobials for treating infections by mycobacteria and other persistent bacteria owing to their potent bactericidal activity against replicating and non-replicating pathogens. However, the clinical utility of rifamycins against Mycobacterium abscessus is seriously compromised by a novel resistance mechanism, namely, rifamycin inactivation by ADP-ribosylation. Using a structure-based approach, we rationally redesign rifamycins through strategic modification of the ansa-chain to block ADP-ribosylation while preserving on-target activity. Validated by a combination of biochemical, structural, and microbiological studies, the most potent analogs overcome ADP-ribosylation, restored their intrinsic low nanomolar activity and demonstrated significant in vivo antibacterial efficacy. Further optimization by tuning drug disposition properties afforded a preclinical candidate with remarkable potency and an outstanding pharmacokinetic profile.
- Department of Medicinal Chemistry, College of Pharmacy, University of Minnesota, 308 SE Harvard St SE, Minneapolis, MN 55455, USA.
Organizational Affiliation: