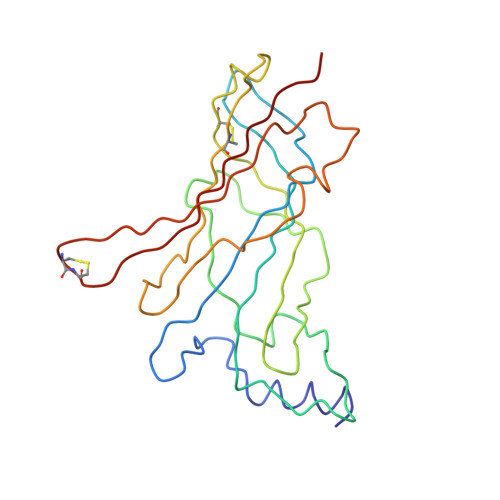
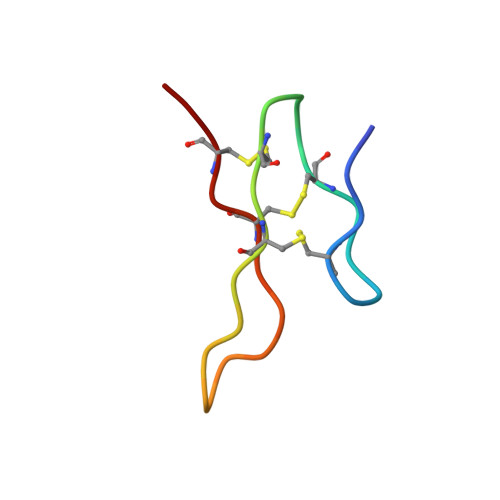
Unravelling the allosteric binding mode of alpha D-VxXXB at nicotinic acetylcholine receptors.
Ho, T.N., Abraham, N., Lewis, R.J.(2023) Front Pharmacol 14: 1170514-1170514
- PubMed: 37124228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fphar.2023.1170514
- Primary Citation Related Structures:
7TXF - PubMed Abstract:
αD-conotoxins are 11 kDa homodimers that potently inhibit nicotinic acetylcholine receptors (nAChRs) through a non-competitive (allosteric) mechanism. In this study, we describe the allosteric binding mode of the granulin-like C-terminal (CTD) of VxXXB bound to Lymnea stagnalis acetylcholine binding protein ( Ls -AChBP), a soluble homologue of the extracellular ligand-binding domain of nAChRs. This co-crystal complex revealed a novel allosteric binding site for nAChR antagonists outside the C-loop that caps the orthosteric site defined by the nAChR agonist nicotine and the antagonist epibatidine. Mutational and docking studies on Ls -AChBP supported a two-site binding mode for full-length VxXXB, with the first CTD binding site located outside the C-loop as seen in the co-crystal complex, with a second CTD binding site located near the N-terminal end of the adjacent subunit of AChBP. These results provide new structural insight into a novel allosteric mechanism of nAChR inhibition and define the cooperative binding mode of the N-terminal domain linked granulin core domains of αD-conotoxins.
- Centre for Pain Research, Institute for Molecular Bioscience, The University of Queensland, St Lucia, QLD, Australia.
Organizational Affiliation: