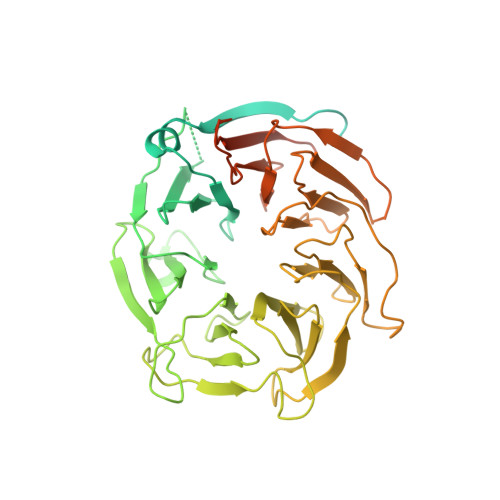
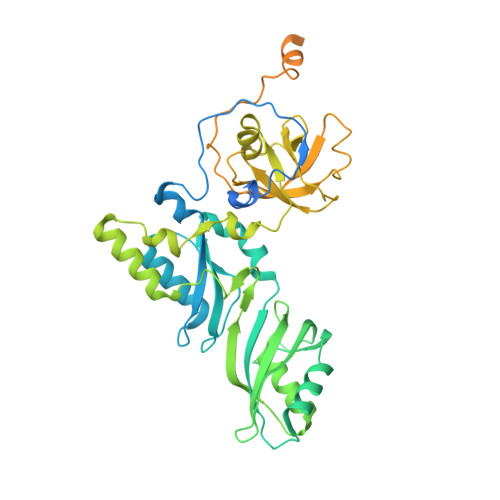
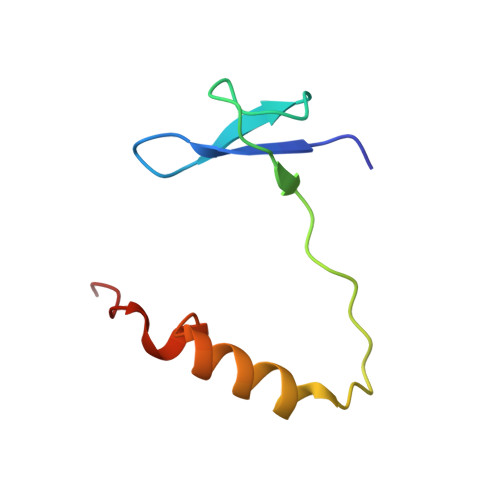
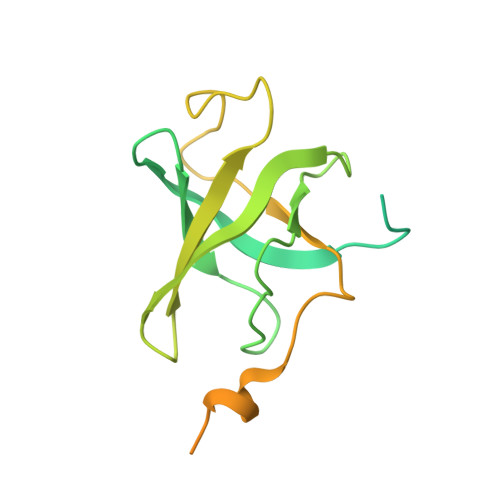
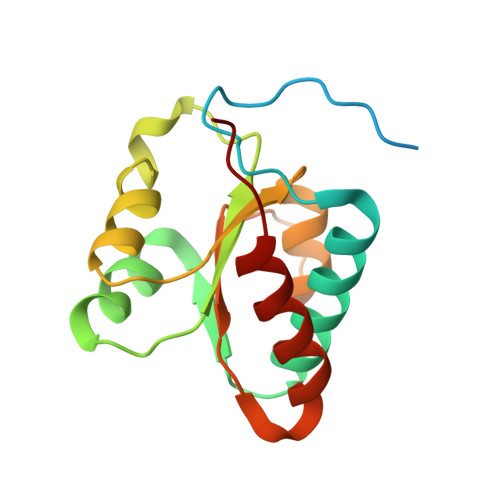
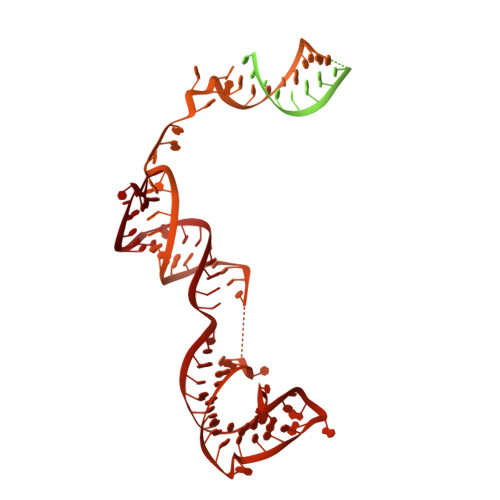
Structure of active human telomerase with telomere shelterin protein TPP1.
Liu, B., He, Y., Wang, Y., Song, H., Zhou, Z.H., Feigon, J.(2022) Nature 604: 578-583
- PubMed: 35418675
- DOI: https://doi.org/10.1038/s41586-022-04582-8
- Primary Citation Related Structures:
7TRC, 7TRD, 7TRE, 7TRF - PubMed Abstract:
Human telomerase is a RNA-protein complex that extends the 3' end of linear chromosomes by synthesizing multiple copies of the telomeric repeat TTAGGG 1 . Its activity is a determinant of cancer progression, stem cell renewal and cellular aging 2-5 . Telomerase is recruited to telomeres and activated for telomere repeat synthesis by the telomere shelterin protein TPP1 6,7 . Human telomerase has a bilobal structure with a catalytic core ribonuclear protein and a H and ACA box ribonuclear protein 8,9 . Here we report cryo-electron microscopy structures of human telomerase catalytic core of telomerase reverse transcriptase (TERT) and telomerase RNA (TER (also known as hTR)), and of telomerase with the shelterin protein TPP1. TPP1 forms a structured interface with the TERT-unique telomerase essential N-terminal domain (TEN) and the telomerase RAP motif (TRAP) that are unique to TERT, and conformational dynamics of TEN-TRAP are damped upon TPP1 binding, defining the requirements for recruitment and activation. The structures further reveal that the elements of TERT and TER that are involved in template and telomeric DNA handling-including the TEN domain and the TRAP-thumb helix channel-are largely structurally homologous to those in Tetrahymena telomerase 10 , and provide unique insights into the mechanism of telomerase activity. The binding site of the telomerase inhibitor BIBR1532 11,12 overlaps a critical interaction between the TER pseudoknot and the TERT thumb domain. Numerous mutations leading to telomeropathies 13,14 are located at the TERT-TER and TEN-TRAP-TPP1 interfaces, highlighting the importance of TER-TERT and TPP1 interactions for telomerase activity, recruitment and as drug targets.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, Los Angeles, CA, USA.
Organizational Affiliation: