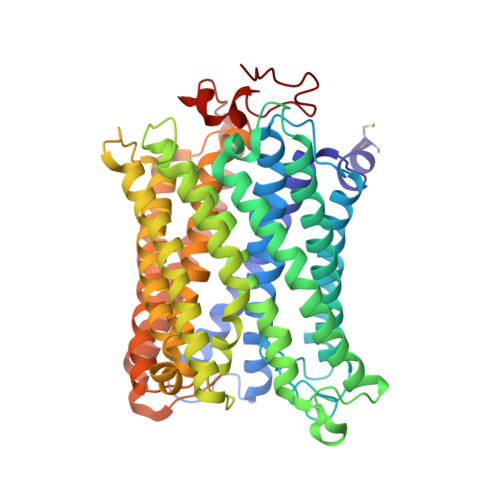
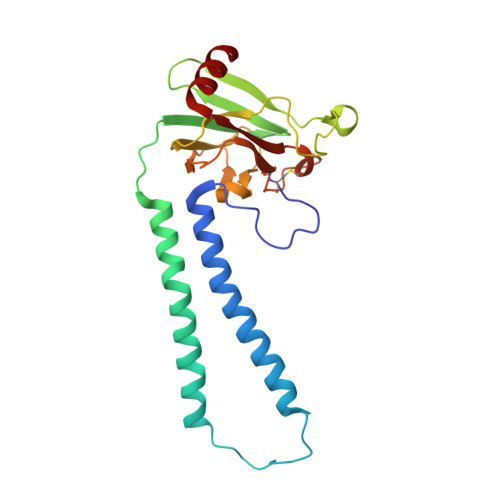
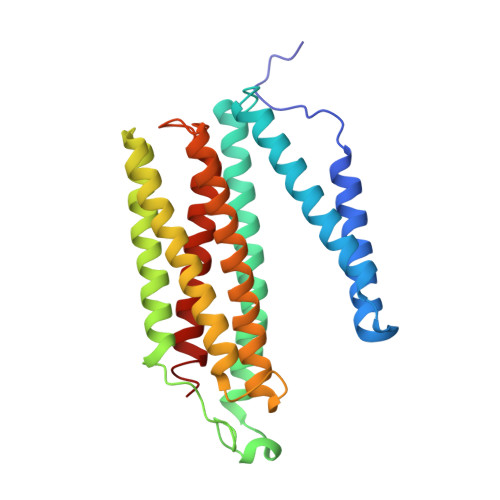
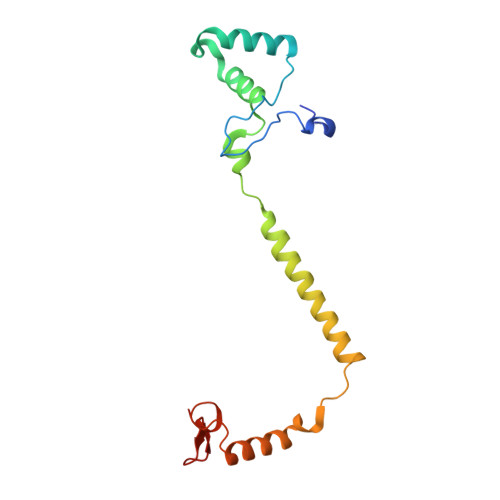
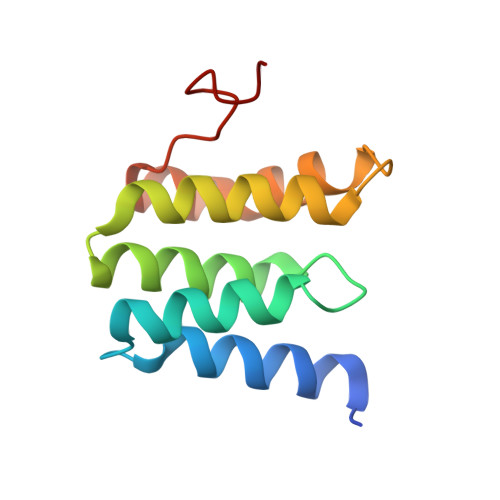
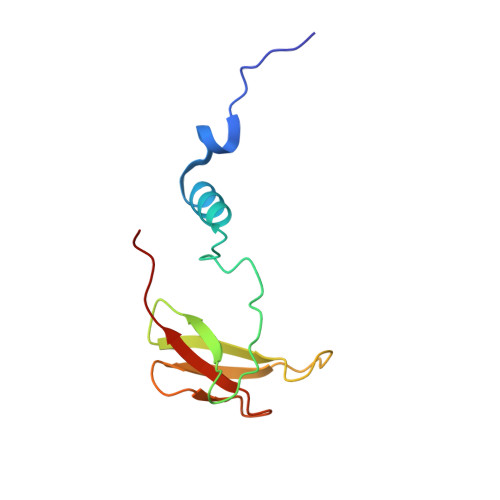
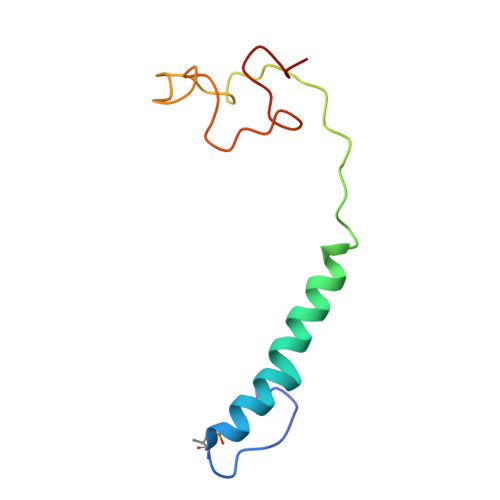
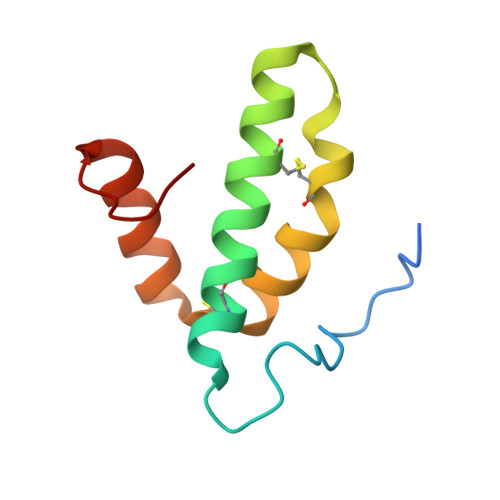
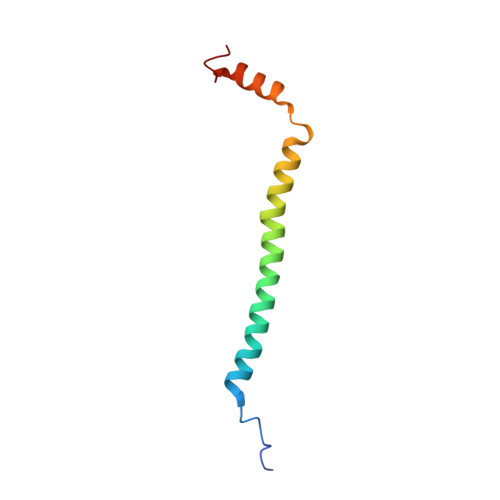
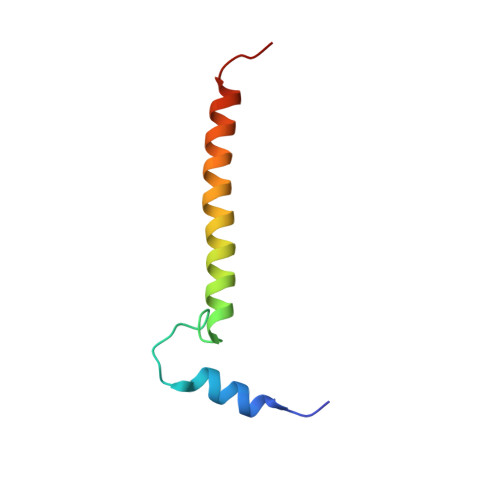
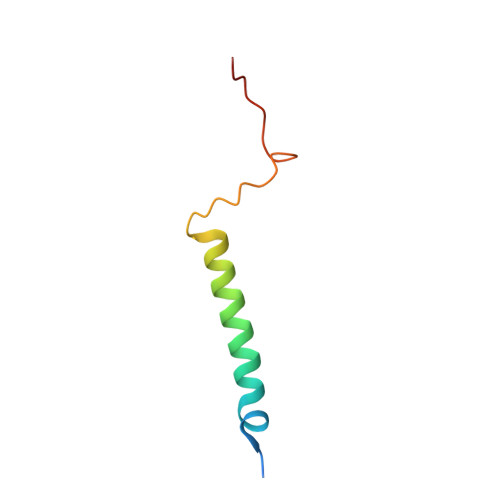
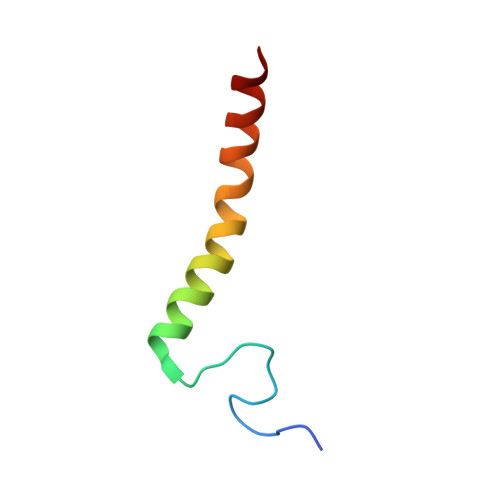
Temperature-dependent structural transition following X-ray-induced metal center reduction in oxidized cytochrome c oxidase.
Ishigami, I., Russi, S., Cohen, A., Yeh, S.R., Rousseau, D.L.(2022) J Biological Chem 298: 101799-101799
- PubMed: 35257742 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.101799
- Primary Citation Related Structures:
7THU, 7TIE, 7TIH, 7TII - PubMed Abstract:
Cytochrome c oxidase (CcO) is the terminal enzyme in the electron transfer chain in the inner membrane of mitochondria. It contains four metal redox centers, two of which, Cu B and heme a 3 , form the binuclear center (BNC), where dioxygen is reduced to water. Crystal structures of CcO in various forms have been reported, from which ligand-binding states of the BNC and conformations of the protein matrix surrounding it have been deduced to elucidate the mechanism by which the oxygen reduction chemistry is coupled to proton translocation. However, metal centers in proteins can be susceptible to X-ray-induced radiation damage, raising questions about the reliability of conclusions drawn from these studies. Here, we used microspectroscopy-coupled X-ray crystallography to interrogate how the structural integrity of bovine CcO in the fully oxidized state (O) is modulated by synchrotron radiation. Spectroscopic data showed that, upon X-ray exposure, O was converted to a hybrid O∗ state where all the four metal centers were reduced, but the protein matrix was trapped in the genuine O conformation and the ligands in the BNC remained intact. Annealing the O∗ crystal above the glass transition temperature induced relaxation of the O∗ structure to a new R∗ structure, wherein the protein matrix converted to the fully reduced R conformation with the exception of helix X, which partly remained in the O conformation because of incomplete dissociation of the ligands from the BNC. We conclude from these data that reevaluation of reported CcO structures obtained with synchrotron light sources is merited.
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York, USA.
Organizational Affiliation: