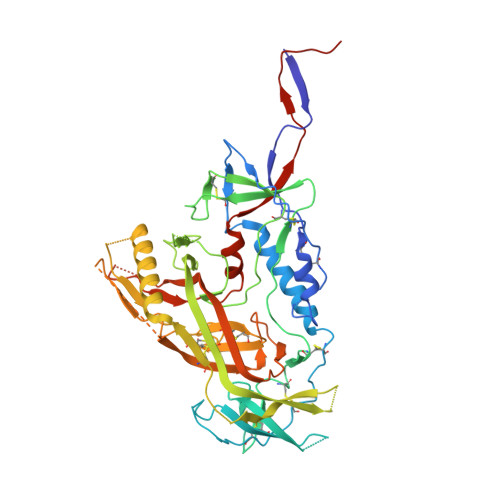
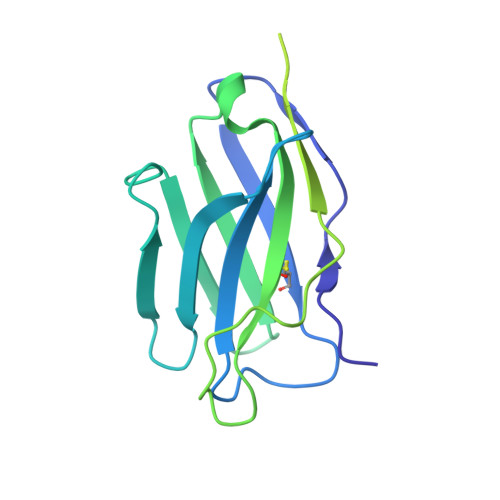
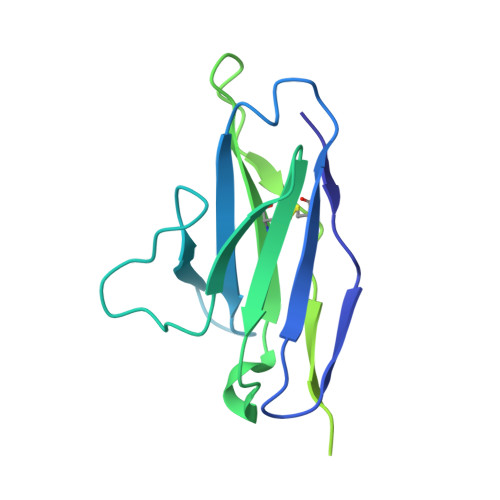
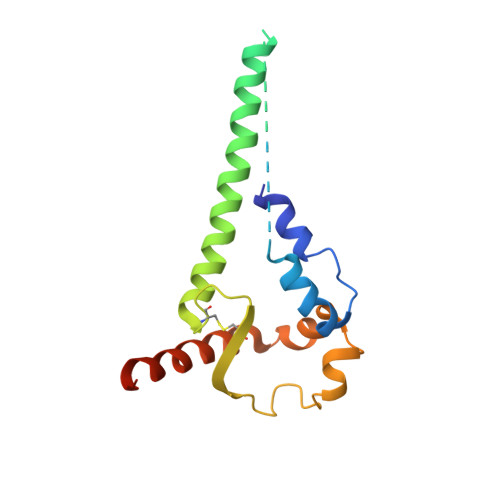
Neutralizing antibodies induced in immunized macaques recognize the CD4-binding site on an occluded-open HIV-1 envelope trimer.
Yang, Z., Dam, K.A., Bridges, M.D., Hoffmann, M.A.G., DeLaitsch, A.T., Gristick, H.B., Escolano, A., Gautam, R., Martin, M.A., Nussenzweig, M.C., Hubbell, W.L., Bjorkman, P.J.(2022) Nat Commun 13: 732-732
- PubMed: 35136084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28424-3
- Primary Citation Related Structures:
7RYU, 7RYV, 7TFN, 7TFO - PubMed Abstract:
Broadly-neutralizing antibodies (bNAbs) against HIV-1 Env can protect from infection. We characterize Ab1303 and Ab1573, heterologously-neutralizing CD4-binding site (CD4bs) antibodies, isolated from sequentially-immunized macaques. Ab1303/Ab1573 binding is observed only when Env trimers are not constrained in the closed, prefusion conformation. Fab-Env cryo-EM structures show that both antibodies recognize the CD4bs on Env trimer with an 'occluded-open' conformation between closed, as targeted by bNAbs, and fully-open, as recognized by CD4. The occluded-open Env trimer conformation includes outwardly-rotated gp120 subunits, but unlike CD4-bound Envs, does not exhibit V1V2 displacement, 4-stranded gp120 bridging sheet, or co-receptor binding site exposure. Inter-protomer distances within trimers measured by double electron-electron resonance spectroscopy suggest an equilibrium between occluded-open and closed Env conformations, consistent with Ab1303/Ab1573 binding stabilizing an existing conformation. Studies of Ab1303/Ab1573 demonstrate that CD4bs neutralizing antibodies that bind open Env trimers can be raised by immunization, thereby informing immunogen design and antibody therapeutic efforts.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA, USA.
Organizational Affiliation: