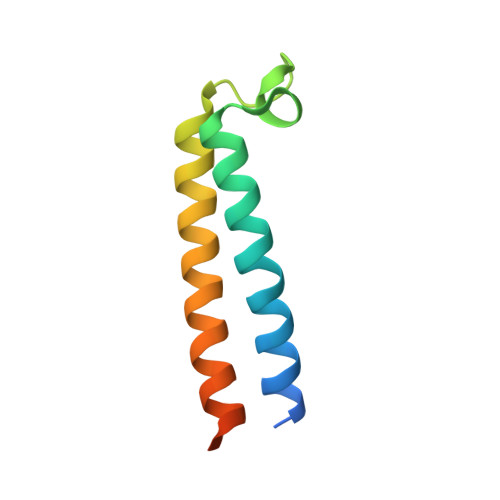
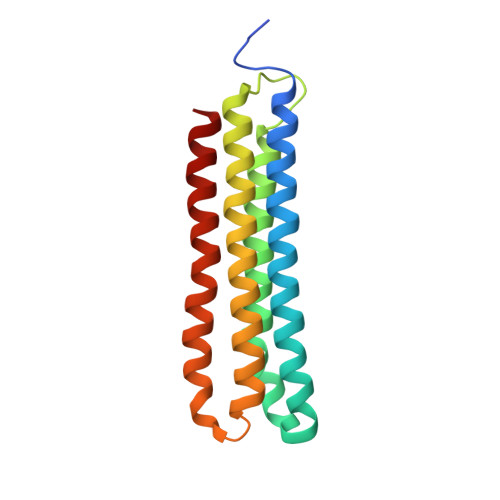
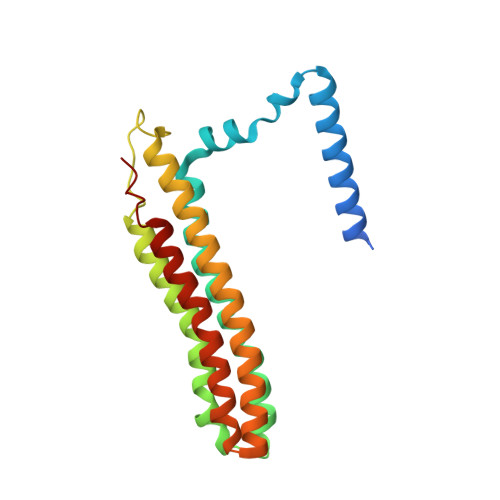
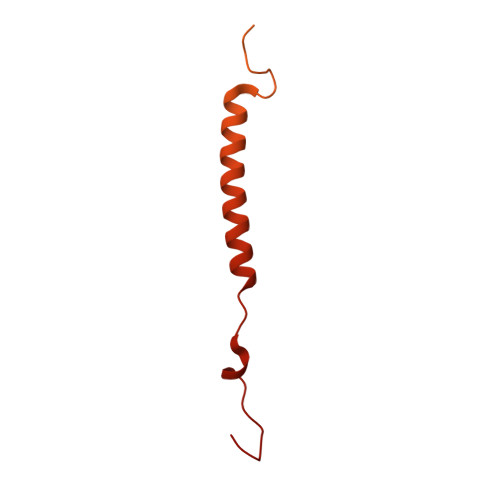
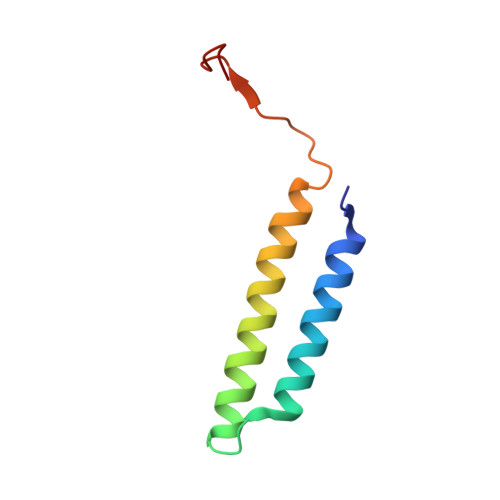
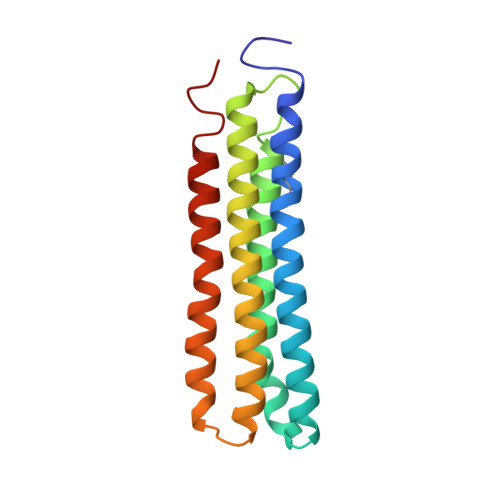
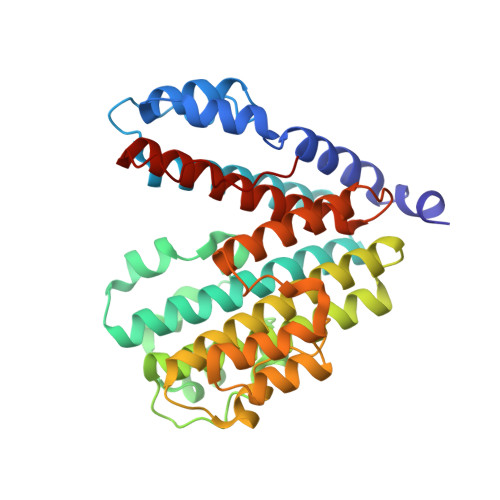
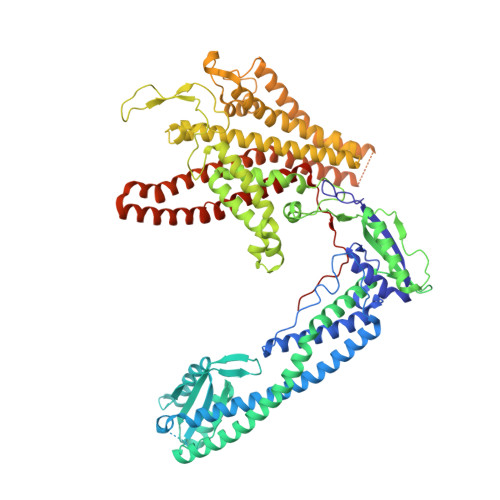
Cryo-EM of the Yeast V O Complex Reveals Distinct Binding Sites for Macrolide V-ATPase Inhibitors.
Keon, K.A., Benlekbir, S., Kirsch, S.H., Muller, R., Rubinstein, J.L.(2022) ACS Chem Biol 17: 619-628
- PubMed: 35148071 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.1c00894
- Primary Citation Related Structures:
7TAO, 7TAP - PubMed Abstract:
Vacuolar-type adenosine triphosphatases (V-ATPases) are proton pumps found in almost all eukaryotic cells. These enzymes consist of a soluble catalytic V 1 region that hydrolyzes ATP and a membrane-embedded V O region responsible for proton translocation. V-ATPase activity leads to acidification of endosomes, phagosomes, lysosomes, secretory vesicles, and the trans-Golgi network, with extracellular acidification occurring in some specialized cells. Small-molecule inhibitors of V-ATPase have played a crucial role in elucidating numerous aspects of cell biology by blocking acidification of intracellular compartments, while therapeutic use of V-ATPase inhibitors has been proposed for the treatment of cancer, osteoporosis, and some infections. Here, we determine structures of the isolated V O complex from Saccharomyces cerevisiae bound to two well-known macrolide inhibitors: bafilomycin A1 and archazolid A. The structures reveal different binding sites for the inhibitors on the surface of the proton-carrying c ring, with only a small amount of overlap between the two sites. Binding of both inhibitors is mediated primarily through van der Waals interactions in shallow pockets and suggests that the inhibitors block rotation of the ring. Together, these structures indicate the existence of a large chemical space available for V-ATPase inhibitors that block acidification by binding the c ring.
- Molecular Medicine Program, The Hospital for Sick Children, Toronto, Canada M5G0A4.
Organizational Affiliation: