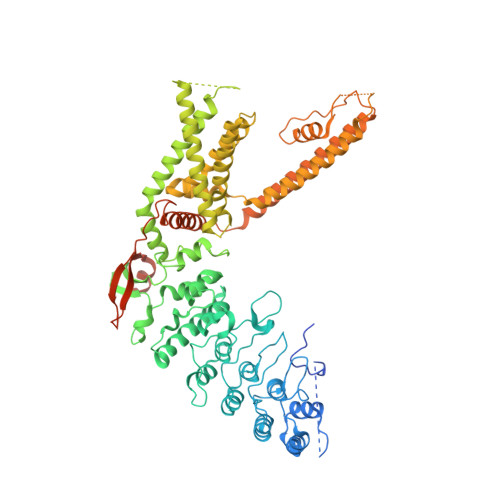
Structural insights into TRPV2 activation by small molecules.
Pumroy, R.A., Protopopova, A.D., Fricke, T.C., Lange, I.U., Haug, F.M., Nguyen, P.T., Gallo, P.N., Sousa, B.B., Bernardes, G.J.L., Yarov-Yarovoy, V., Leffler, A., Moiseenkova-Bell, V.Y.(2022) Nat Commun 13: 2334-2334
- PubMed: 35484159 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30083-3
- Primary Citation Related Structures:
7N0M, 7N0N, 7T37, 7T38 - PubMed Abstract:
Transient receptor potential vanilloid 2 (TRPV2) is involved in many critical physiological and pathophysiological processes, making it a promising drug target. Here we present cryo-electron microscopy (cryo-EM) structures of rat TRPV2 in lipid nanodiscs activated by 2-aminoethoxydiphenyl borate (2-APB) and propose a TRPV2-specific 2-ABP binding site at the interface of S5 of one monomer and the S4-S5 linker of the adjacent monomer. In silico docking and electrophysiological studies confirm the key role of His521 and Arg539 in 2-APB activation of TRPV2. Additionally, electrophysiological experiments show that the combination of 2-APB and cannabidiol has a synergetic effect on TRPV2 activation, and cryo-EM structures demonstrate that both drugs were able to bind simultaneously. Together, our cryo-EM structures represent multiple functional states of the channel, providing a native picture of TRPV2 activation by small molecules and a structural framework for the development of TRPV2-specific activators.
- Department of Systems Pharmacology and Translational Therapeutics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, 19104, United States.
Organizational Affiliation: