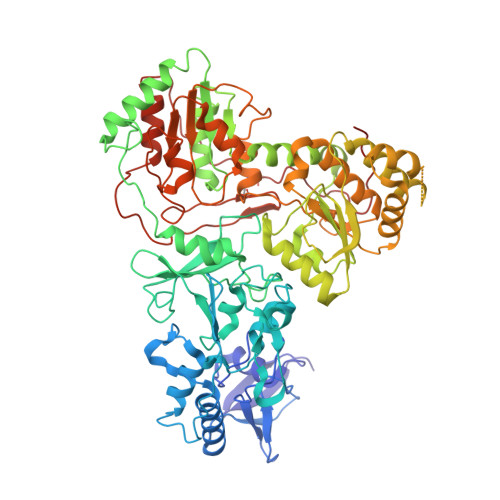
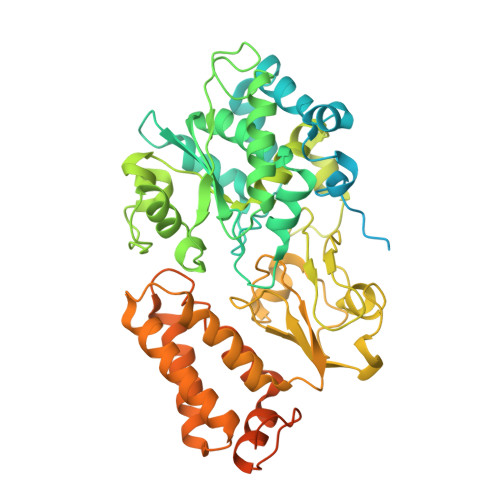
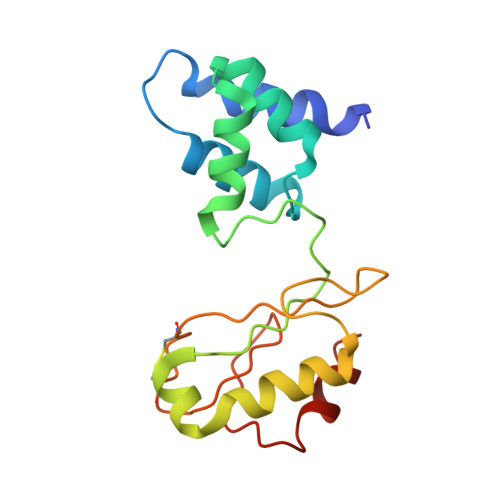
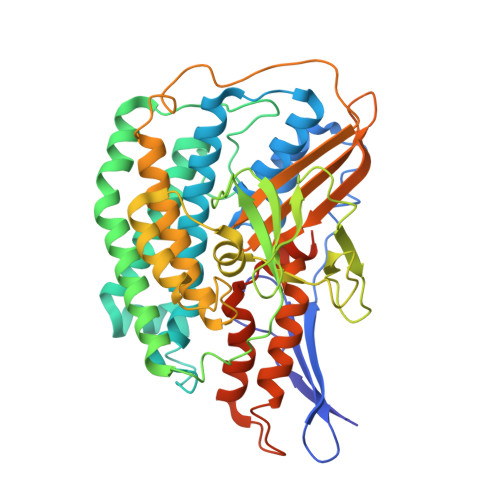
Structure and electron transfer pathways of an electron-bifurcating NiFe-hydrogenase.
Feng, X., Schut, G.J., Haja, D.K., Adams, M.W.W., Li, H.(2022) Sci Adv 8: eabm7546-eabm7546
- PubMed: 35213221 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abm7546
- Primary Citation Related Structures:
7T2R, 7T30 - PubMed Abstract:
Electron bifurcation enables thermodynamically unfavorable biochemical reactions. Four groups of bifurcating flavoenzyme are known and three use FAD to bifurcate. FeFe-HydABC hydrogenase represents the fourth group, but its bifurcation site is unknown. We report cryo-EM structures of the related NiFe-HydABCSL hydrogenase that reversibly oxidizes H 2 and couples endergonic reduction of ferredoxin with exergonic reduction of NAD. FMN surrounded by a unique arrangement of iron sulfur clusters forms the bifurcating center. NAD binds to FMN in HydB, and electrons from H 2 via HydA to a HydB [4Fe-4S] cluster enable the FMN to reduce NAD. Low-potential electron transfer from FMN to the HydC [2Fe-2S] cluster and subsequent reduction of a uniquely penta-coordinated HydB [2Fe-2S] cluster require conformational changes, leading to ferredoxin binding and reduction by a [4Fe-4S] cluster in HydB. This work clarifies the electron transfer pathways for a large group of hydrogenases underlying many essential functions in anaerobic microorganisms.
- Department of Structural Biology, Van Andel Institute, Grand Rapids, MI, USA.
Organizational Affiliation: