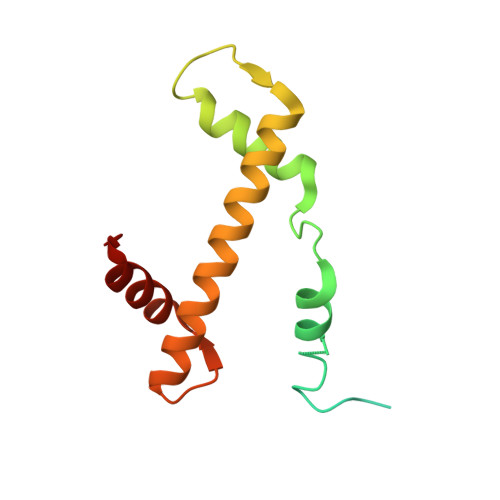
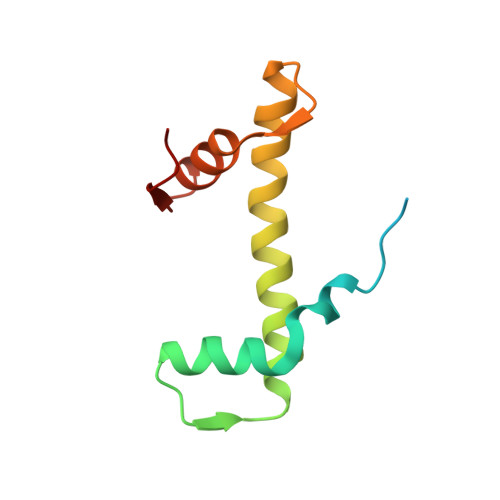
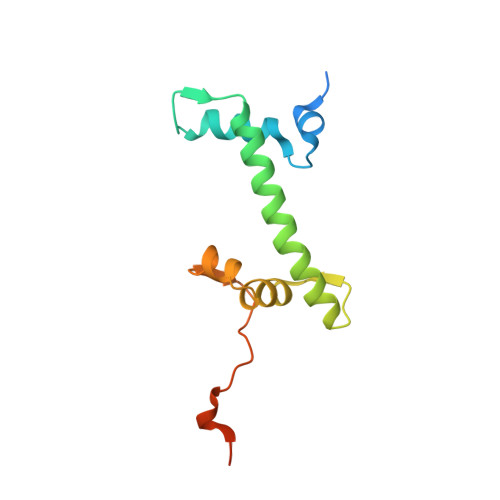
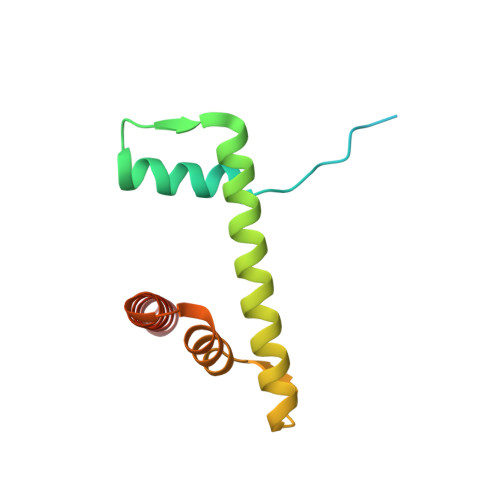
Nucleosome recognition and DNA distortion by the Chd1 remodeler in a nucleotide-free state.
Nodelman, I.M., Das, S., Faustino, A.M., Fried, S.D., Bowman, G.D., Armache, J.P.(2022) Nat Struct Mol Biol 29: 121-129
- PubMed: 35173352 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00719-x
- Primary Citation Related Structures:
7SWY, 7TN2 - PubMed Abstract:
Chromatin remodelers are ATP-dependent enzymes that reorganize nucleosomes within all eukaryotic genomes. Here we report a complex of the Chd1 remodeler bound to a nucleosome in a nucleotide-free state, determined by cryo-EM to 2.3 Å resolution. The remodeler stimulates the nucleosome to absorb an additional nucleotide on each strand at two different locations: on the tracking strand within the ATPase binding site and on the guide strand one helical turn from the ATPase motor. Remarkably, the additional nucleotide on the tracking strand is associated with a local transformation toward an A-form geometry, explaining how sequential ratcheting of each DNA strand occurs. The structure also reveals a histone-binding motif, ChEx, which can block opposing remodelers on the nucleosome and may allow Chd1 to participate in histone reorganization during transcription.
- Thomas C. Jenkins Department of Biophysics, Johns Hopkins University, Baltimore, MD, USA.
Organizational Affiliation: