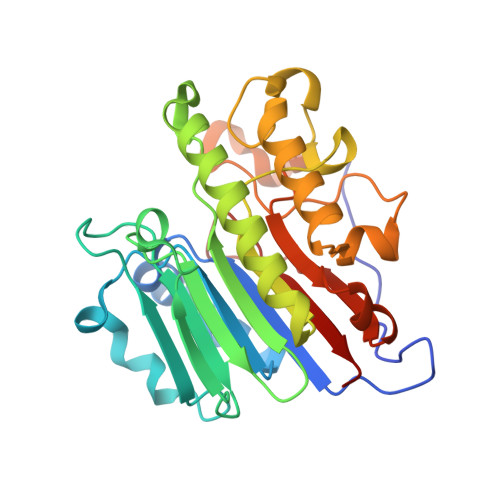
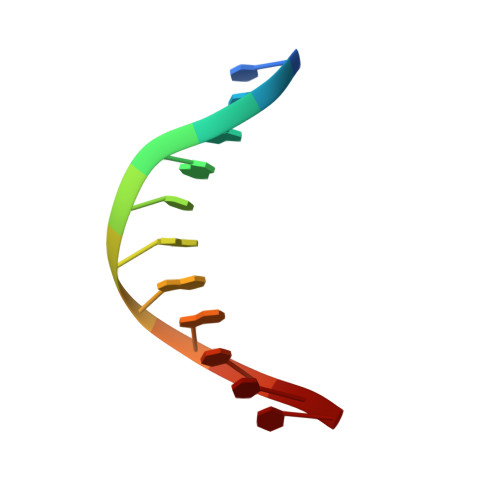
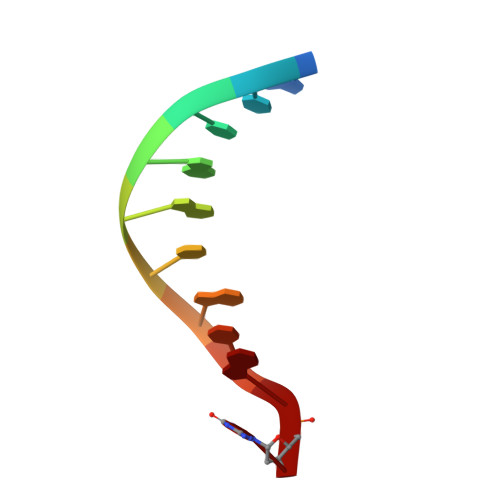
Processing oxidatively damaged bases at DNA strand breaks by APE1.
Whitaker, A.M., Stark, W.J., Freudenthal, B.D.(2022) Nucleic Acids Res 50: 9521-9533
- PubMed: 36018803 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac695
- Primary Citation Related Structures:
7SUV, 7SVB - PubMed Abstract:
Reactive oxygen species attack the structure of DNA, thus altering its base-pairing properties. Consequently, oxidative stress-associated DNA lesions are a major source of the mutation load that gives rise to cancer and other diseases. Base excision repair (BER) is the pathway primarily tasked with repairing DNA base damage, with apurinic/apyrimidinic endonuclease (APE1) having both AP-endonuclease and 3' to 5' exonuclease (exo) DNA cleavage functions. The lesion 8-oxo-7,8-dihydroguanine (8-oxoG) can enter the genome as either a product of direct damage to the DNA, or through polymerase insertion at the 3'-end of a DNA strand during replication or repair. Importantly, 3'-8-oxoG impairs the ligation step of BER and therefore must be removed by the exo activity of a surrogate enzyme to prevent double stranded breaks and cell death. In the present study, we use X-ray crystallography to characterize the exo activity of APE1 on 3'-8-oxoG substrates. These structures support a unified APE1 exo mechanism that differs from its more canonical AP-endonuclease activity. In addition, through complementation of the structural data with enzyme kinetics and binding studies employing both wild-type and rationally designed APE1 mutants, we were able to identify and characterize unique protein: DNA contacts that specifically mediate 8-oxoG removal by APE1.
- Department of Biochemistry and Molecular Biology, University of Kansas Medical Center, Kansas City, KS 66160, USA.
Organizational Affiliation: