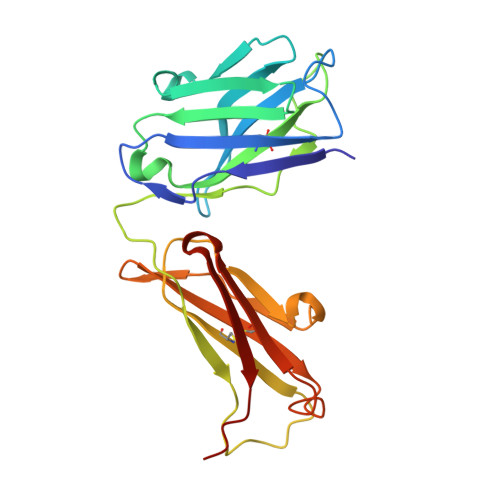
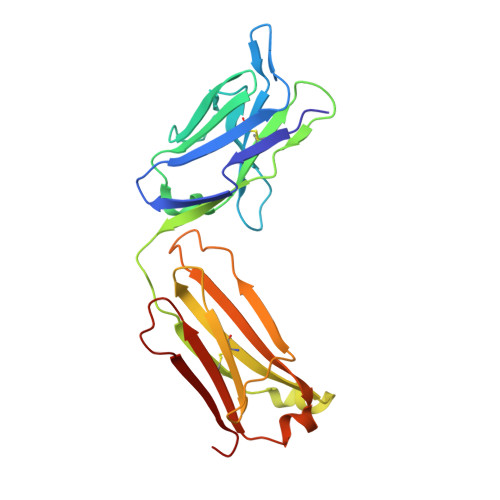
Monoclonal antibody 7H2.2 binds the C-terminus of the cancer-oocyte antigen SAS1B through the hydrophilic face of a conserved amphipathic helix corresponding to one of only two regions predicted to be ordered
Legg, M.S.G., Gagnon, S.M.L., Powell, C.J., Boulanger, M.J., Li, A.J.J., Evans, S.V.(2022) Acta Crystallogr D Biol Crystallogr 78: 623-632
- PubMed: 35503210 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798322003011
- Primary Citation Related Structures:
7ST8 - PubMed Abstract:
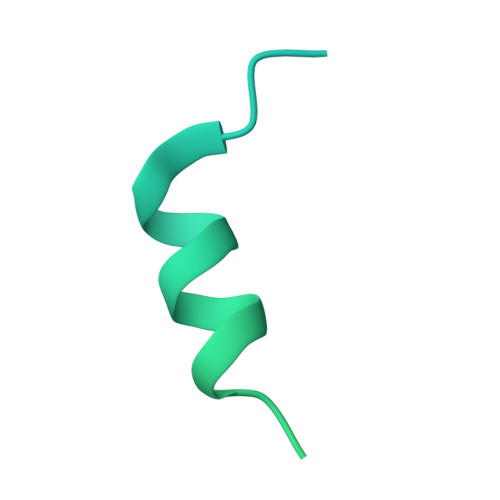
The structure of the antigen-binding fragment (Fab) of mouse monoclonal antibody 7H2.2 in complex with a 15-residue fragment from the metalloproteinase sperm acrosomal SLLP1 binding protein (SAS1B), which is a molecular and cellular candidate for both cancer therapy and female contraception, has been determined at 2.75 Å resolution by single-crystal X-ray diffraction. Although the crystallization conditions contained the final 148 C-terminal residues of SAS1B, the Fab was observed to crystallize in complex with a 15-residue fragment corresponding to one of only two elements of secondary structure that are predicted to be ordered within the C-terminal region of SAS1B. The antigen forms an amphipathic α-helix that binds the 7H2.2 combining site via hydrophilic residues in an epitope that spans the length of the antigen α-helix, with only two CH-π interactions observed along the edge of the interface between the antibody and antigen. Interestingly, the paratope contains two residues mutated away from the germline (YL32F and YH58R), as well as a ProH96-ThrH97-AspH98-AspH99 insertion within heavy chain CDR3. The intact 7H2.2 antibody exhibits high affinity for the SAS1B antigen, with 1:1 binding and nanomolar affinity for both the SAS1B C-terminal construct used for crystallization (3.38 ± 0.59 nM) and a 15-amino-acid synthetic peptide construct corresponding to the helical antigen observed within the crystal structure (1.60 ± 0.31 nM). The SAS1B-antibody structure provides the first structural insight into any portion of the subdomain architecture of the C-terminal region of the novel cancer-oocyte tumor surface neoantigen SAS1B and provides a basis for the targeted use of SAS1B.
- Department of Biochemistry and Microbiology, University of Victoria, PO Box 3055 STN CSC, Victoria, BC V8Z 7X8, Canada.
Organizational Affiliation: