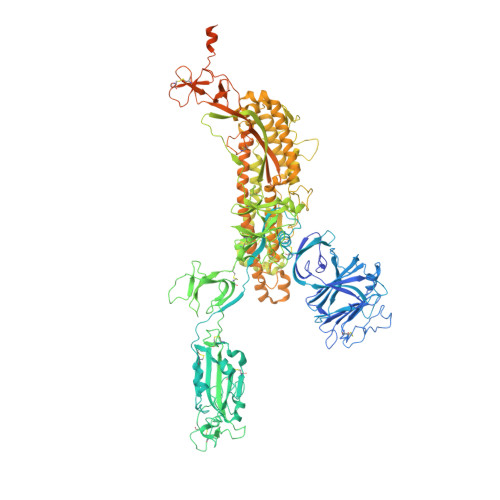
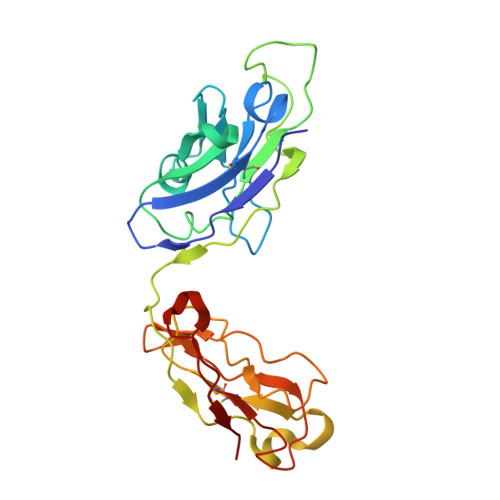
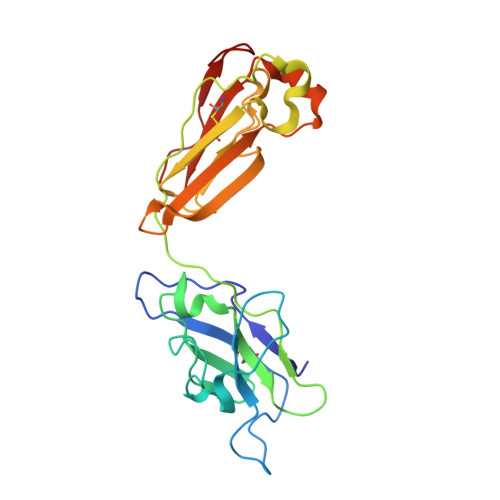
A broadly cross-reactive antibody neutralizes and protects against sarbecovirus challenge in mice.
Martinez, D.R., Schafer, A., Gobeil, S., Li, D., De la Cruz, G., Parks, R., Lu, X., Barr, M., Stalls, V., Janowska, K., Beaudoin, E., Manne, K., Mansouri, K., Edwards, R.J., Cronin, K., Yount, B., Anasti, K., Montgomery, S.A., Tang, J., Golding, H., Shen, S., Zhou, T., Kwong, P.D., Graham, B.S., Mascola, J.R., Montefiori, D.C., Alam, S.M., Sempowski, G., Sempowski, G.D., Khurana, S., Wiehe, K., Saunders, K.O., Acharya, P., Haynes, B.F., Baric, R.S.(2022) Sci Transl Med 14: eabj7125-eabj7125
- PubMed: 34726473 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scitranslmed.abj7125
- Primary Citation Related Structures:
7SG4 - PubMed Abstract:
Severe acute respiratory syndrome coronaviruses 1 (SARS-CoV) and 2 (SARS-CoV-2), including SARS-CoV-2 variants of concern, can cause deadly infections. The mortality associated with sarbecovirus infection underscores the importance of developing broadly effective countermeasures against them, which could be key in the prevention and mitigation of current and future zoonotic events. Here, we demonstrate the neutralization of SARS-CoV; bat coronaviruses WIV-1 and RsSHC014; and SARS-CoV-2 variants D614G, B.1.1.7, B.1.351, P.1, B.1.429, B.1.526, B.1.617.1, and B.1.617.2 by a receptor binding domain (RBD)–specific human antibody, DH1047. Prophylactic and therapeutic treatment with DH1047 was protective against SARS-CoV, WIV-1, RsSHC014, and SARS-CoV-2 B.1.351 infection in mice. Binding and structural analysis showed high affinity binding of DH1047 to an epitope that is highly conserved among sarbecoviruses. Thus, DH1047 is a broadly protective antibody that can prevent infection and mitigate outbreaks caused by SARS-related strains and SARS-CoV-2 variants. Our results also suggest that the conserved RBD epitope bound by DH1047 is a rational target for a universal sarbecovirus vaccine.
- Department of Epidemiology, University of North Carolina at Chapel Hill, Chapel Hill, NC, USA.
Organizational Affiliation: