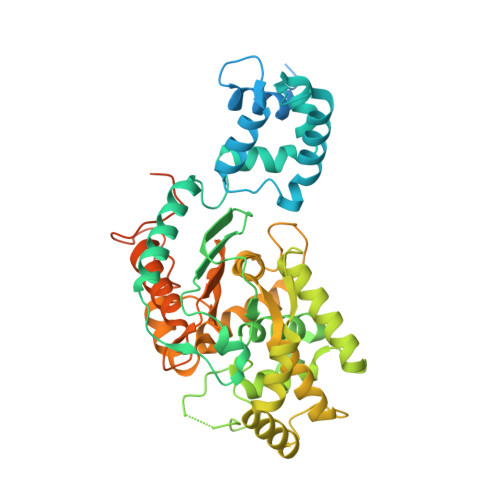
High-resolution structures of mitochondrial glutaminase C tetramers indicate conformational changes upon phosphate binding.
Nguyen, T.T., Ramachandran, S., Hill, M.J., Cerione, R.A.(2022) J Biological Chem 298: 101564-101564
- PubMed: 34999118 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.101564
- Primary Citation Related Structures:
7SBM, 7SBN - PubMed Abstract:
The mitochondrial enzyme glutaminase C (GAC) is upregulated in many cancer cells to catalyze the first step in glutamine metabolism, the hydrolysis of glutamine to glutamate. The dependence of cancer cells on this transformed metabolic pathway highlights GAC as a potentially important therapeutic target. GAC acquires maximal catalytic activity upon binding to anionic activators such as inorganic phosphate. To delineate the mechanism of GAC activation, we used the tryptophan substitution of tyrosine 466 in the catalytic site of the enzyme as a fluorescent reporter for glutamine binding in the presence and absence of phosphate. We show that in the absence of phosphate, glutamine binding to the Y466W GAC tetramer exhibits positive cooperativity. A high-resolution X-ray structure of tetrameric Y466W GAC bound to glutamine suggests that cooperativity in substrate binding is coupled to tyrosine 249, located at the edge of the catalytic site (i.e., the "lid"), adopting two distinct conformations. In one dimer within the GAC tetramer, the lids are open and glutamine binds weakly, whereas, in the adjoining dimer, the lids are closed over the substrates, resulting in higher affinity interactions. When crystallized in the presence of glutamine and phosphate, all four subunits of the Y466W GAC tetramer exhibited bound glutamine with closed lids. Glutamine can bind with high affinity to each subunit, which subsequently undergo simultaneous catalysis. These findings explain how the regulated transitioning of GAC between different conformational states ensures that maximal catalytic activity is reached in cancer cells only when an allosteric activator is available.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York, USA.
Organizational Affiliation: