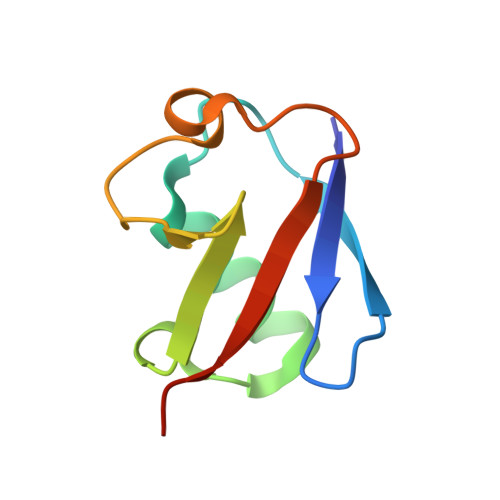
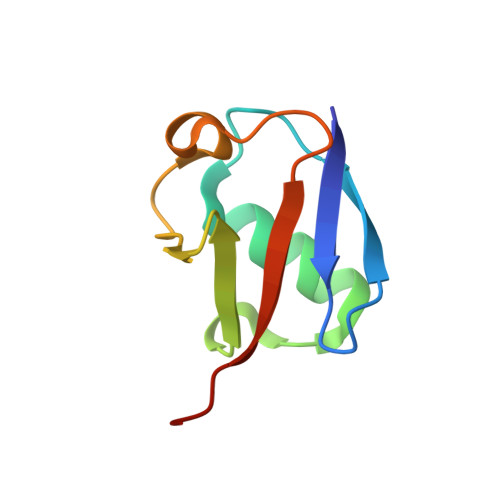
Dual domain recognition determines SARS-CoV-2 PLpro selectivity for human ISG15 and K48-linked di-ubiquitin.
Wydorski, P.M., Osipiuk, J., Lanham, B.T., Tesar, C., Endres, M., Engle, E., Jedrzejczak, R., Mullapudi, V., Michalska, K., Fidelis, K., Fushman, D., Joachimiak, A., Joachimiak, L.A.(2023) Nat Commun 14: 2366-2366
- PubMed: 37185902 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38031-5
- Primary Citation Related Structures:
7RBR, 7RBS, 7S6O, 7S6P, 7UV5 - PubMed Abstract:
The Papain-like protease (PLpro) is a domain of a multi-functional, non-structural protein 3 of coronaviruses. PLpro cleaves viral polyproteins and posttranslational conjugates with poly-ubiquitin and protective ISG15, composed of two ubiquitin-like (UBL) domains. Across coronaviruses, PLpro showed divergent selectivity for recognition and cleavage of posttranslational conjugates despite sequence conservation. We show that SARS-CoV-2 PLpro binds human ISG15 and K48-linked di-ubiquitin (K48-Ub 2 ) with nanomolar affinity and detect alternate weaker-binding modes. Crystal structures of untethered PLpro complexes with ISG15 and K48-Ub 2 combined with solution NMR and cross-linking mass spectrometry revealed how the two domains of ISG15 or K48-Ub 2 are differently utilized in interactions with PLpro. Analysis of protein interface energetics predicted differential binding stabilities of the two UBL/Ub domains that were validated experimentally. We emphasize how substrate recognition can be tuned to cleave specifically ISG15 or K48-Ub 2 modifications while retaining capacity to cleave mono-Ub conjugates. These results highlight alternative druggable surfaces that would inhibit PLpro function.
- Molecular Biophysics Graduate Program, University of Texas Southwestern Medical Center, Dallas, TX, 75390, USA.
Organizational Affiliation: