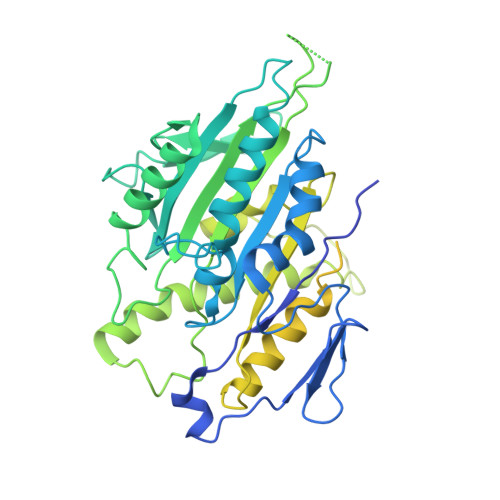
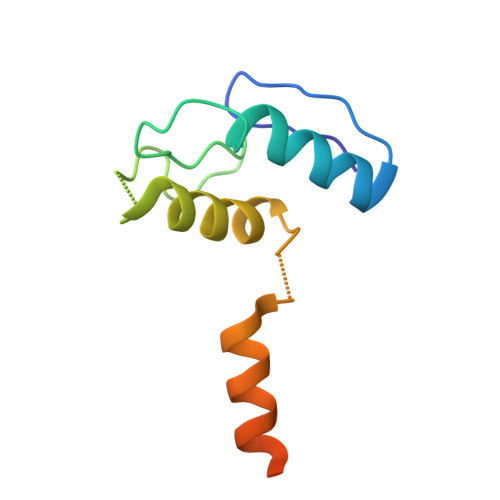
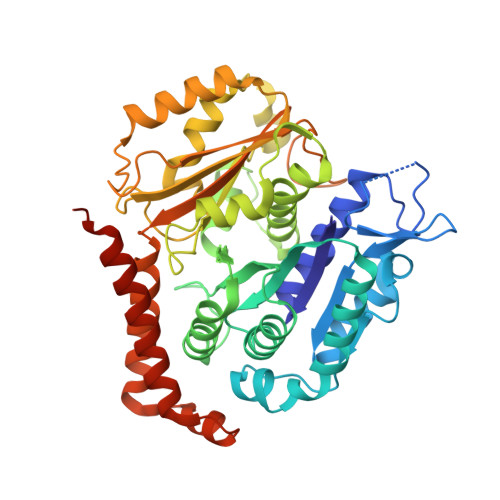
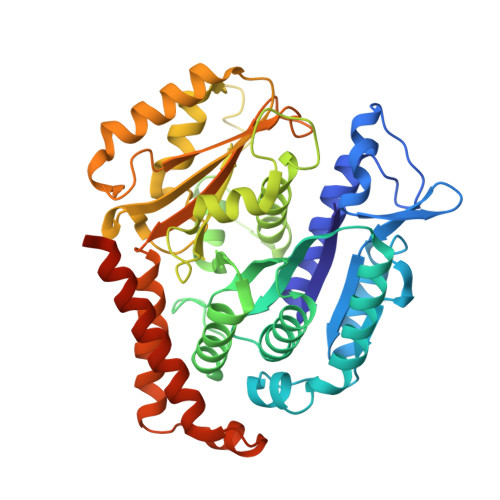
Cytoskeletal regulation of a transcription factor by DNA mimicry via coiled-coil interactions.
Haque, F., Freniere, C., Ye, Q., Mani, N., Wilson-Kubalek, E.M., Ku, P.I., Milligan, R.A., Subramanian, R.(2022) Nat Cell Biol 24: 1088-1098
- PubMed: 35725768
- DOI: https://doi.org/10.1038/s41556-022-00935-7
- Primary Citation of Related Structures:
7RX0 - PubMed Abstract:
A long-established strategy for transcription regulation is the tethering of transcription factors to cellular membranes. By contrast, the principal effectors of Hedgehog signalling, the GLI transcription factors, are regulated by microtubules in the primary cilium and the cytoplasm. How GLI is tethered to microtubules remains unclear. Here, we uncover DNA mimicry by the ciliary kinesin KIF7 as a mechanism for the recruitment of GLI to microtubules, wherein the coiled-coil dimerization domain of KIF7, characterized by its striking shape, size and charge similarity to DNA, forms a complex with the DNA-binding zinc fingers in GLI, thus revealing a mode of tethering a DNA-binding protein to the cytoskeleton. GLI increases KIF7 microtubule affinity and consequently modulates the localization of both proteins to microtubules and the cilium tip. Thus, the kinesin-microtubule system is not a passive GLI tether but a regulatable platform tuned by the kinesin-transcription factor interaction. We retooled this coiled-coil-based GLI-KIF7 interaction to inhibit the nuclear and cilium localization of GLI. This strategy can potentially be exploited to downregulate erroneously activated GLI in human cancers.
- Department of Molecular Biology, Massachusetts General Hospital, Boston, MA, USA.
Organizational Affiliation: