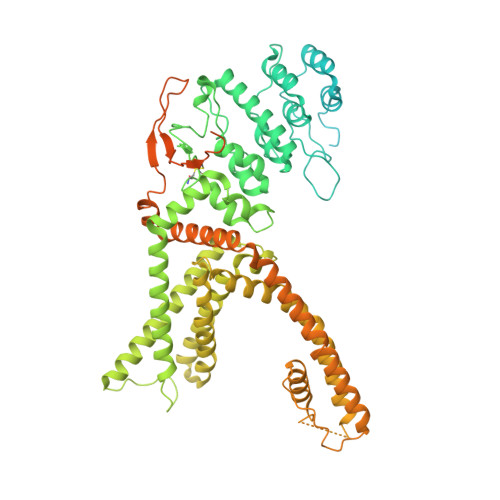
Vanilloid-dependent TRPV1 opening trajectory from cryoEM ensemble analysis.
Kwon, D.H., Zhang, F., Fedor, J.G., Suo, Y., Lee, S.Y.(2022) Nat Commun 13: 2874-2874
- PubMed: 35610228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30602-2
- Primary Citation Related Structures:
7RQU, 7RQV, 7RQW, 7RQX, 7RQY, 7RQZ - PubMed Abstract:
Single particle cryo-EM often yields multiple protein conformations within a single dataset, but experimentally deducing the temporal relationship of these conformers within a conformational trajectory is not trivial. Here, we use thermal titration methods and cryo-EM in an attempt to obtain temporal resolution of the conformational trajectory of the vanilloid receptor TRPV1 with resiniferatoxin (RTx) bound. Based on our cryo-EM ensemble analysis, RTx binding to TRPV1 appears to induce intracellular gate opening first, followed by selectivity filter dilation, then pore loop rearrangement to reach the final open state. This apparent conformational wave likely arises from the concerted, stepwise, additive structural changes of TRPV1 over many subdomains. Greater understanding of the RTx-mediated long-range allostery of TRPV1 could help further the therapeutic potential of RTx, which is a promising drug candidate for pain relief associated with advanced cancer or knee arthritis.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC, 27710, USA.
Organizational Affiliation: