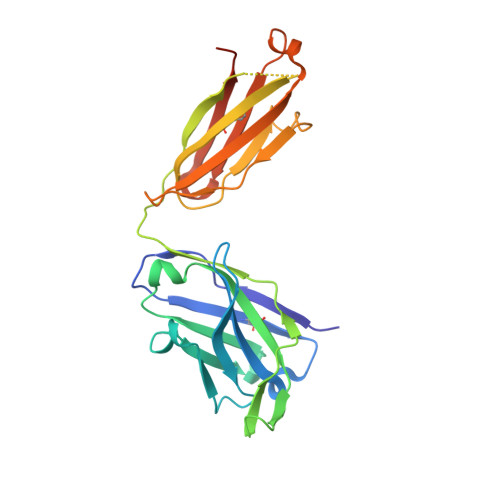
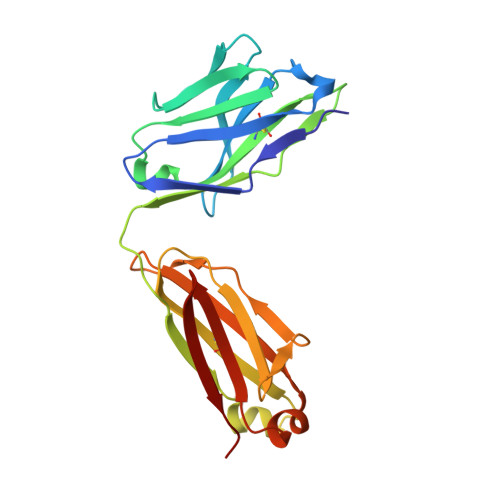
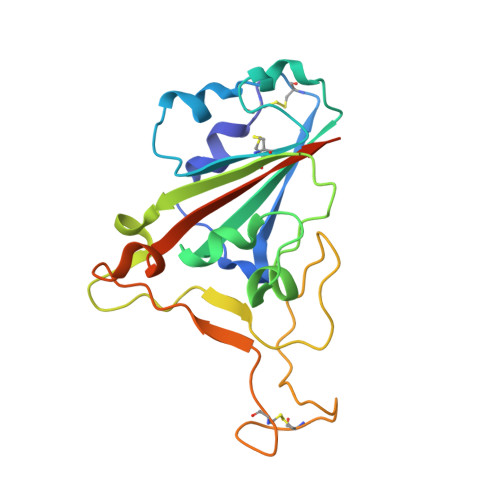
Broad cross-reactivity across sarbecoviruses exhibited by a subset of COVID-19 donor-derived neutralizing antibodies.
Jette, C.A., Cohen, A.A., Gnanapragasam, P.N.P., Muecksch, F., Lee, Y.E., Huey-Tubman, K.E., Schmidt, F., Hatziioannou, T., Bieniasz, P.D., Nussenzweig, M.C., West Jr., A.P., Keeffe, J.R., Bjorkman, P.J., Barnes, C.O.(2021) Cell Rep 36: 109760-109760
- PubMed: 34534459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2021.109760
- Primary Citation Related Structures:
7RKS, 7RKU, 7RKV - PubMed Abstract:
Many anti-severe acute respiratory syndrome coronavirus 2 (anti-SARS-CoV-2) neutralizing antibodies target the angiotensin-converting enzyme 2 (ACE2) binding site on viral spike receptor-binding domains (RBDs). Potent antibodies recognize exposed variable epitopes, often rendering them ineffective against other sarbecoviruses and SARS-CoV-2 variants. Class 4 anti-RBD antibodies against a less-exposed, but more-conserved, cryptic epitope could recognize newly emergent zoonotic sarbecoviruses and variants, but they usually show only weak neutralization potencies. Here, we characterize two class 4 anti-RBD antibodies derived from coronavirus disease 2019 (COVID-19) donors that exhibit breadth and potent neutralization of zoonotic coronaviruses and SARS-CoV-2 variants. C118-RBD and C022-RBD structures reveal orientations that extend from the cryptic epitope to occlude ACE2 binding and CDRH3-RBD main-chain H-bond interactions that extend an RBD β sheet, thus reducing sensitivity to RBD side-chain changes. A C118-spike trimer structure reveals rotated RBDs that allow access to the cryptic epitope and the potential for intra-spike crosslinking to increase avidity. These studies facilitate vaccine design and illustrate potential advantages of class 4 RBD-binding antibody therapeutics.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: