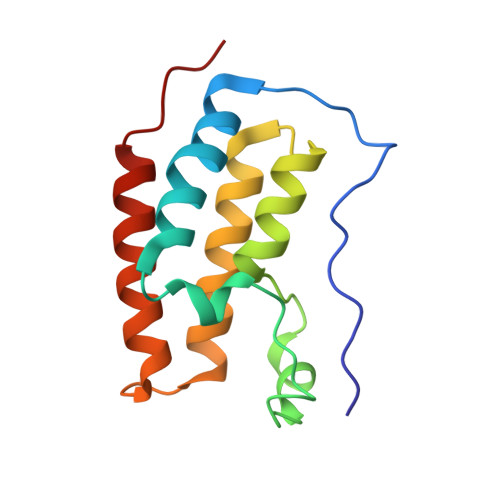
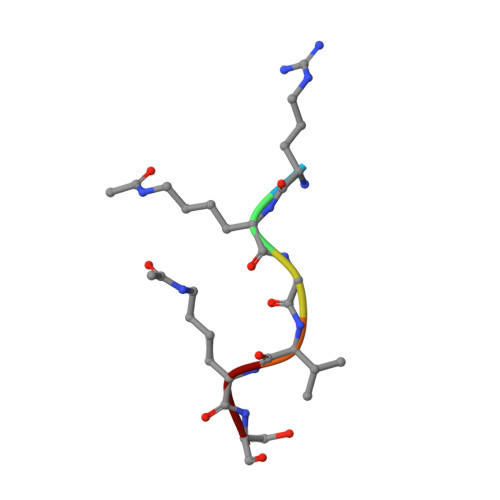
Uncovering the Bromodomain Interactome using Site-Specific Azide-Acetyllysine Photochemistry, Proteomic Profiling and Structural Characterization
Wagner, S., Fedorov, E., Sudhamalla, B., Jnawali, H.N., Debiec, R., Ghosh, A., Islam, K.To be published.