Characterization of the stepwise engineering and optimization of a retargeted DNA binding protein and gene-editing meganuclease
Werther, R.A., Ubilla-Rodriguez, N.C., Smiley, A.T., Havens, K., Lambert, A.R., Stoddard, B.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| I-OnuI_e-hPD1-e | 300 | synthetic construct | Mutation(s): 0 |  | |
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (26-MER) | 26 | synthetic construct |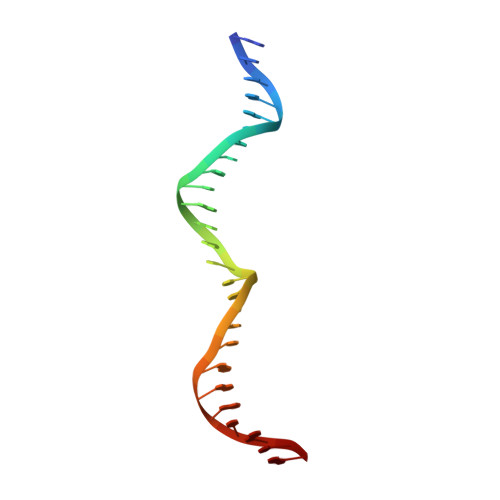  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (26-MER) | 26 | synthetic construct |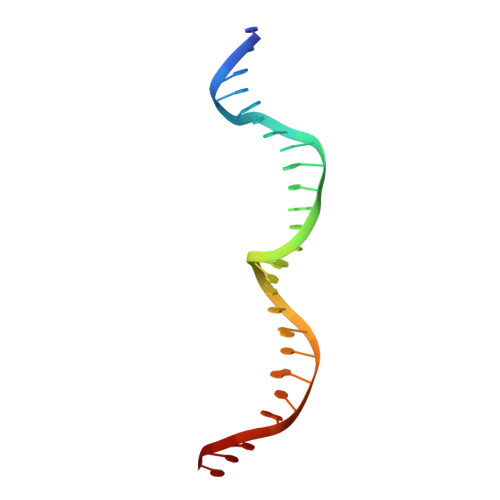  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EDO Download:Ideal Coordinates CCD File | D [auth A], E [auth A], K [auth B] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| CA Download:Ideal Coordinates CCD File | F [auth A], G [auth A], H [auth A] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | I [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Download:Ideal Coordinates CCD File | J [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 41.55 | α = 90 |
| b = 64.055 | β = 90 |
| c = 171.296 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| HKL-2000 | data processing |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | NIH R01 GM105691 |
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | NIH R01 GM 139752 |
| Other private | United States | Funding from bluebird bio |