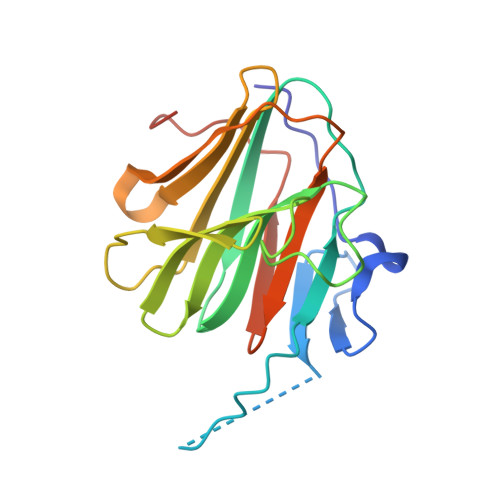
Structural analysis of TRIM family PRYSPRY domains and its implications for E3-ligand design.
Zhubi, R., Chaikuad, A., Munoz Sosa, C.J., Joerger, A.C., Knapp, S.(2025) J Struct Biol X 12: 100134-100134
- PubMed: 40821731
- DOI: https://doi.org/10.1016/j.yjsbx.2025.100134
- Primary Citation of Related Structures:
7B2S, 7QRY, 7QRZ, 7QS0, 7QS1, 7QS2, 7QS3, 7QS4, 7QS5, 9R11 - PubMed Abstract:
Tripartite motif (TRIM) proteins constitute one of the largest subfamilies of RING-type E3 ubiquitin ligases and are attractive targets for the development of novel degraders that exploit the ubiquitin-proteasome pathway. More than half of all TRIM family members contain a PRYSPRY domain, a potentially druggable protein interaction module, located in their C-terminal region. Here, we have determined crystal structures of the PRYSPRY domains from nine TRIM family proteins: TRIM1 (MID2), TRIM9, TRIM10, TRIM11, TRIM15, TRIM16, TRIM18 (MID1), TRIM36, and TRIM67. These structures reveal conservation of the overall β-sandwich topology, despite low sequence conservation, with a unique subdomain swap observed in TRIM11. Significant variations were found in the loops flanking the canonical substrate-binding site, which modulate the shape and electrostatic properties of the binding pocket, hinting at substantial differences in substrate specificity and binding modes among family members. TRIM36 features a unique structural motif between the canonical β-strands 2 and 3, leading to the formation of a dimer, with the canonical substrate-binding site partially occluded by the dimerization motif. In addition, we mapped the locations of missense mutations in MID1 associated with X-linked Opitz syndrome, suggesting that some of these mutations impair the conformational stability of the protein. Taken together, our data provide intriguing insights into the structural and functional divergence of TRIM family PRYSPRY domains, their potential druggability and substrate recognition, and the challenges of ligand design.
- Institute of Pharmaceutical Chemistry, Goethe University, Max-von-Laue-Str. 9, 60438 Frankfurt am Main, Germany.
Organizational Affiliation: