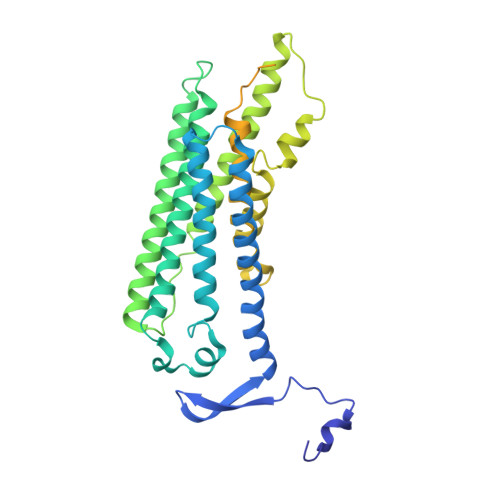
Activation mechanism of the class D fungal GPCR dimer Ste2.
Velazhahan, V., Ma, N., Vaidehi, N., Tate, C.G.(2022) Nature 603: 743-748
- PubMed: 35296853
- DOI: https://doi.org/10.1038/s41586-022-04498-3
- Primary Citation of Related Structures:
7QA8, 7QB9, 7QBC, 7QBI - PubMed Abstract:
The fungal class D1 G-protein-coupled receptor (GPCR) Ste2 has a different arrangement of transmembrane helices compared with mammalian GPCRs and a distinct mode of coupling to the heterotrimeric G protein Gpa1-Ste2-Ste18 1 . In addition, Ste2 lacks conserved sequence motifs such as DRY, PIF and NPXXY, which are associated with the activation of class A GPCRs 2 . This suggested that the activation mechanism of Ste2 may also differ. Here we determined structures of Saccharomyces cerevisiae Ste2 in the absence of G protein in two different conformations bound to the native agonist α-factor, bound to an antagonist and without ligand. These structures revealed that Ste2 is indeed activated differently from other GPCRs. In the inactive state, the cytoplasmic end of transmembrane helix H7 is unstructured and packs between helices H1-H6, blocking the G protein coupling site. Agonist binding results in the outward movement of the extracellular ends of H6 and H7 by 6 Å. On the intracellular surface, the G protein coupling site is formed by a 20 Å outward movement of the unstructured region in H7 that unblocks the site, and a 12 Å inward movement of H6. This is a distinct mechanism in GPCRs, in which the movement of H6 and H7 upon agonist binding facilitates G protein coupling.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, UK.
Organizational Affiliation: