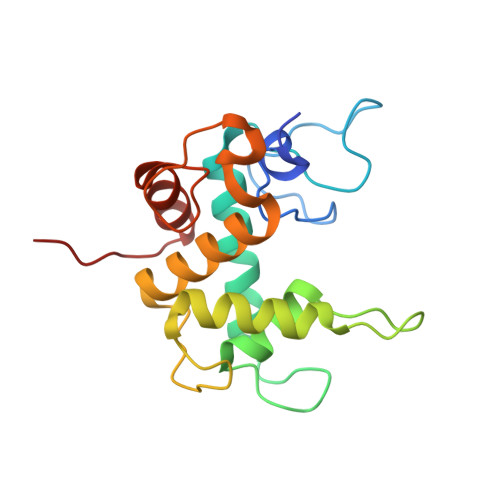
Diversity of the lysozyme fold: structure of the catalytic domain from an unusual endolysin encoded by phage Enc34.
Cernooka, E., Rumnieks, J., Zrelovs, N., Tars, K., Kazaks, A.(2022) Sci Rep 12: 5005-5005
- PubMed: 35322067 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-022-08765-1
- Primary Citation Related Structures:
7Q47 - PubMed Abstract:
Endolysins are bacteriophage-encoded peptidoglycan-degrading enzymes with potential applications for treatment of multidrug-resistant bacterial infections. Hafnia phage Enc34 encodes an unusual endolysin with an N-terminal enzymatically active domain and a C-terminal transmembrane domain. The catalytic domain of the endolysin belongs to the conserved protein family PHA02564 which has no recognizable sequence similarity to other known endolysin types. Turbidity reduction assays indicate that the Enc34 enzyme is active against peptidoglycan from a variety of Gram-negative bacteria including the opportunistic pathogen Pseudomonas aeruginosa PAO1. The crystal structure of the catalytic domain of the Enc34 endolysin shows a distinctive all-helical architecture that distantly resembles the α-lobe of the lysozyme fold. Conserved catalytically important residues suggest a shared evolutionary history between the Enc34 endolysin and GH73 and GH23 family glycoside hydrolases and propose a molecular signature for substrate cleavage for a large group of peptidoglycan-degrading enzymes.
- Latvian Biomedical Research and Study Centre, Ratsupites 1 k-1, Riga, 1067, Latvia.
Organizational Affiliation: