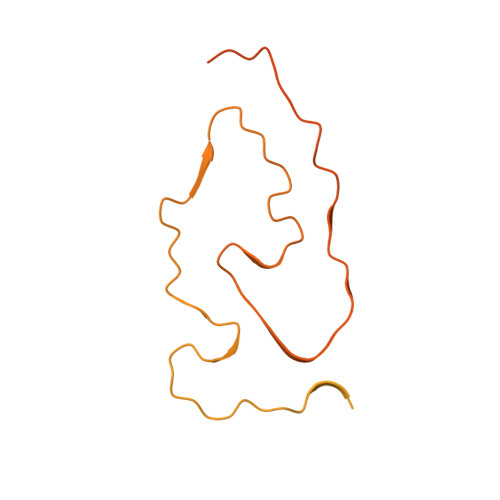
Structure of pathological TDP-43 filaments from ALS with FTLD.
Arseni, D., Hasegawa, M., Murzin, A.G., Kametani, F., Arai, M., Yoshida, M., Ryskeldi-Falcon, B.(2022) Nature 601: 139-143
- PubMed: 34880495 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-04199-3
- Primary Citation Related Structures:
7PY2 - PubMed Abstract:
The abnormal aggregation of TAR DNA-binding protein 43 kDa (TDP-43) in neurons and glia is the defining pathological hallmark of the neurodegenerative disease amyotrophic lateral sclerosis (ALS) and multiple forms of frontotemporal lobar degeneration (FTLD) 1,2 . It is also common in other diseases, including Alzheimer's and Parkinson's. No disease-modifying therapies exist for these conditions and early diagnosis is not possible. The structures of pathological TDP-43 aggregates are unknown. Here we used cryo-electron microscopy to determine the structures of aggregated TDP-43 in the frontal and motor cortices of an individual who had ALS with FTLD and from the frontal cortex of a second individual with the same diagnosis. An identical amyloid-like filament structure comprising a single protofilament was found in both brain regions and individuals. The ordered filament core spans residues 282-360 in the TDP-43 low-complexity domain and adopts a previously undescribed double-spiral-shaped fold, which shows no similarity to those of TDP-43 filaments formed in vitro 3,4 . An abundance of glycine and neutral polar residues facilitates numerous turns and restricts β-strand length, which results in an absence of β-sheet stacking that is associated with cross-β amyloid structure. An uneven distribution of residues gives rise to structurally and chemically distinct surfaces that face external densities and suggest possible ligand-binding sites. This work enhances our understanding of the molecular pathogenesis of ALS and FTLD and informs the development of diagnostic and therapeutic agents that target aggregated TDP-43.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: