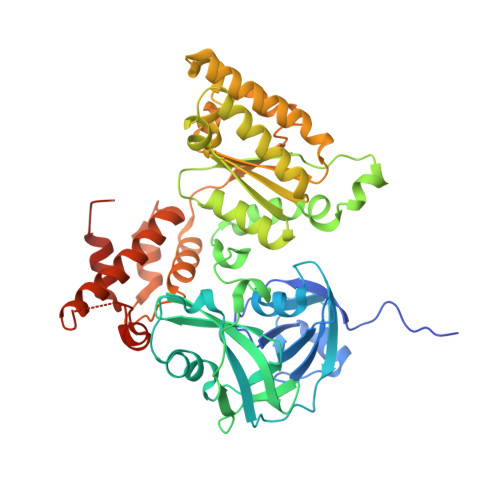
Fragment screening using biolayer interferometry reveals ligands targeting the SHP-motif binding site of the AAA+ ATPase p97
Bothe, S., Hanzelmann, P., Bohler, S., Kehrein, J., Zehe, M., Wiedemann, C., Hellmich, U.A., Brenk, R., Schindelin, H., Sotriffer, C.(2022) Commun Chem 5: 169