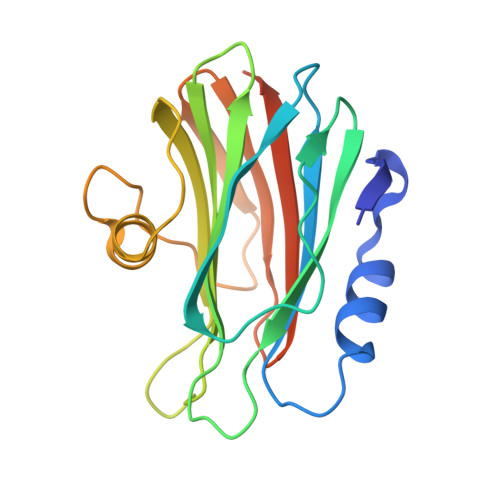
Pore-forming moss protein bryoporin is structurally and mechanistically related to actinoporins from evolutionarily distant cnidarians.
Solinc, G., Svigelj, T., Omersa, N., Snoj, T., Pirc, K., Znidarsic, N., Yamaji-Hasegawa, A., Kobayashi, T., Anderluh, G., Podobnik, M.(2022) J Biological Chem 298: 102455-102455
- PubMed: 36063994 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102455
- Primary Citation Related Structures:
7PUD - PubMed Abstract:
Pore-forming proteins perforate lipid membranes and consequently affect their integrity and cell fitness. Therefore, it is not surprising that many of these proteins from bacteria, fungi, or certain animals act as toxins. While pore-forming proteins have also been found in plants, there is little information about their molecular structure and mode of action. Bryoporin is a protein from the moss Physcomitrium patens, and its corresponding gene was found to be upregulated by various abiotic stresses, especially dehydration, as well as upon fungal infection. Based on the amino acid sequence, it was suggested that bryoporin was related to the actinoporin family of pore-forming proteins, originally discovered in sea anemones. Here, we provide the first detailed structural and functional analysis of this plant cytolysin. The crystal structure of monomeric bryoporin is highly similar to those of actinoporins. Our cryo-EM analysis of its pores showed an actinoporin-like octameric structure, thereby revealing a close kinship of proteins from evolutionarily distant organisms. This was further confirmed by our observation of bryoporin's preferential binding to and formation of pores in membranes containing animal sphingolipids, such as sphingomyelin and ceramide phosphoethanolamine; however, its binding affinity was weaker than that of actinoporin equinatoxin II. We determined bryoporin did not bind to major sphingolipids found in fungi or plants, and its membrane-binding and pore-forming activity was enhanced by various sterols. Our results suggest that bryoporin could represent a part of the moss defense arsenal, acting as a pore-forming toxin against membranes of potential animal pathogens, parasites, or predators.
- Department of Molecular Biology and Nanobiotechnology, National Institute of Chemistry, Ljubljana, Slovenia.
Organizational Affiliation: