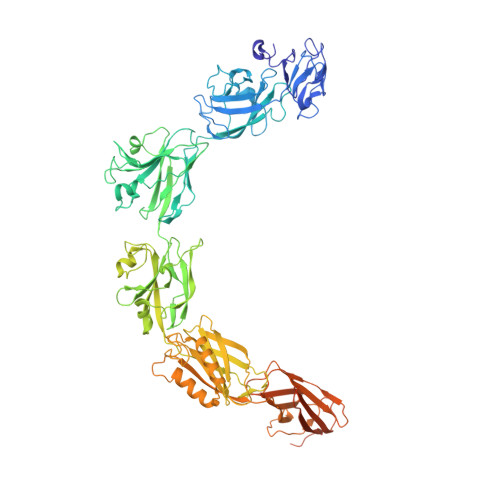
Complete atomic structure of a native archaeal cell surface.
von Kugelgen, A., Alva, V., Bharat, T.A.M.(2021) Cell Rep 37: 110052-110052
- PubMed: 34818541
- DOI: https://doi.org/10.1016/j.celrep.2021.110052
- Primary Citation Related Structures:
7PTP, 7PTR, 7PTT, 7PTU - PubMed Abstract:
Many prokaryotic cells are covered by an ordered, proteinaceous, sheet-like structure called a surface layer (S-layer). S-layer proteins (SLPs) are usually the highest copy number macromolecules in prokaryotes, playing critical roles in cellular physiology such as blocking predators, scaffolding membranes, and facilitating environmental interactions. Using electron cryomicroscopy of two-dimensional sheets, we report the atomic structure of the S-layer from the archaeal model organism Haloferax volcanii. This S-layer consists of a hexagonal array of tightly interacting immunoglobulin-like domains, which are also found in SLPs across several classes of archaea. Cellular tomography reveal that the S-layer is nearly continuous on the cell surface, completed by pentameric defects in the hexagonal lattice. We further report the atomic structure of the SLP pentamer, which shows markedly different relative arrangements of SLP domains needed to complete the S-layer. Our structural data provide a framework for understanding cell surfaces of archaea at the atomic level.
- Sir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, UK.
Organizational Affiliation: