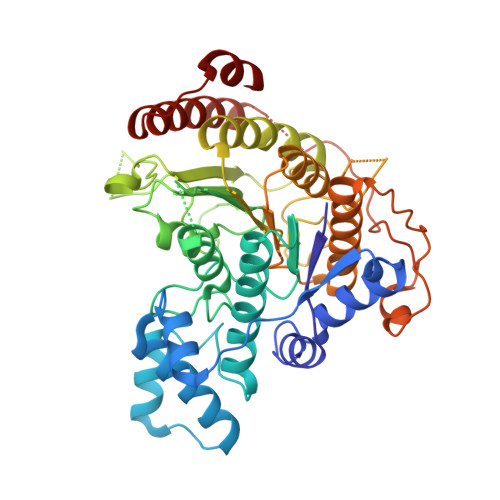
Crystal structures of Schistosoma mansoni histone deacetylase 8 reveal a novel binding site for allosteric inhibitors.
Saccoccia, F., Pozzetti, L., Gimmelli, R., Butini, S., Guidi, A., Papoff, G., Giannaccari, M., Brogi, S., Scognamiglio, V., Gemma, S., Ruberti, G., Campiani, G.(2022) J Biological Chem 298: 102375-102375
- PubMed: 35970392
- DOI: https://doi.org/10.1016/j.jbc.2022.102375
- Primary Citation Related Structures:
7P2S, 7P2T, 7P2U, 7P2V, 7POZ - PubMed Abstract:
Parasitic diseases cause significant global morbidity and mortality particularly in the poorest regions of the world. Schistosomiasis, one of the most widespread neglected tropical diseases, affects more than 200 million people worldwide. Histone deacetylase (HDAC) inhibitors are prominent epigenetic drugs that are being investigated in the treatment of several diseases, including cancers and parasitic diseases. Schistosoma mansoni HDAC8 (SmHDAC8) is highly expressed in all life cycle stages of the parasite, and selective inhibition is required in order to avoid undesirable off-target effects in the host. Herein, by X-ray crystal structures of SmHDAC8-inhibitor complexes, biochemical and phenotypic studies, we found two schistosomicidal spiroindoline derivatives binding a novel site, next to Trp198, on the enzyme surface. We determined that by acting on this site, either by mutation of the Trp198 or by compound binding, a decrease in the activity of the enzyme is achieved. Remarkably, this allosteric site differs from the human counterpart; rather, it is conserved in all Schistosoma species, as well as Rhabidoptera and Trematoda classes, thus paving the way for the design of HDAC8-selective allosteric inhibitors with improved properties.
- Institute of Biochemistry and Cell Biology, Italian National Research Council (IBBC-CNR), Adriano Buzzati-Traverso Campus, Monterotondo Scalo, Rome, Italy. Electronic address: fulvio.saccoccia@cnr.it.
Organizational Affiliation: