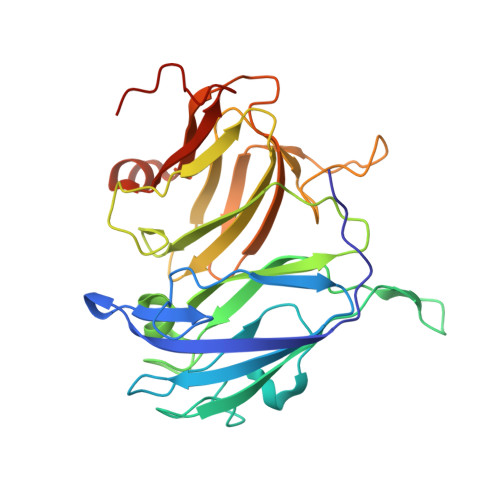
Engineering the Catalytic Properties of Two-Domain Laccase from Streptomyces griseoflavus Ac-993.
Kolyadenko, I., Scherbakova, A., Kovalev, K., Gabdulkhakov, A., Tishchenko, S.(2021) Int J Mol Sci 23
- PubMed: 35008493
- DOI: https://doi.org/10.3390/ijms23010065
- Primary Citation Related Structures:
7PEN, 7PES, 7PFR, 7PTM, 7PU0, 7PUH - PubMed Abstract:
Laccases catalyze the oxidation of substrates with the concomitant reduction of oxygen to water. Recently, we found that polar residues located in tunnels leading to Cu2 and Cu3 ions control oxygen entrance (His 165) and proton transport (Arg 240) of two-domain laccase (2D) from Streptomyces griseoflavus (SgfSL). In this work, we have focused on optimizing the substrate-binding pocket (SBP) of SgfSL while simultaneously adjusting the oxygen reduction process. SgfSL variants with three single (Met199Ala, Met199Gly, and Tyr230Ala) and three double amino acid residues substitutions (Met199Gly/His165Ala, His165Ala/Arg240His, Met199Gly/Arg240His) were constructed, purified, and investigated. Combination of substitutions in the SBP and in the tunnel leading to Cu2 ion (Met199Gly/Arg240His) increased SgfSL catalytic activity towards ABTS by 5-fold, and towards 2.6-DMP by 16-fold. The high activity of the Met199Gly/Arg240His variant can be explained by the combined effect of the SBP geometry optimization (Met199Gly) and increased proton flux via the tunnel leading to Cu2 ion (Arg240His). Moreover, the variant with Met199Gly and His165Ala mutations did not significantly increase SgfSL's activity, but led to a drastic shift in the optimal pH of 2.6-DMP oxidation. These results indicate that His 165 not only regulates oxygen access, but it also participates in proton transport in 2D laccases.
- Institute of Protein Research RAS, 142290 Pushchino, Russia.
Organizational Affiliation: