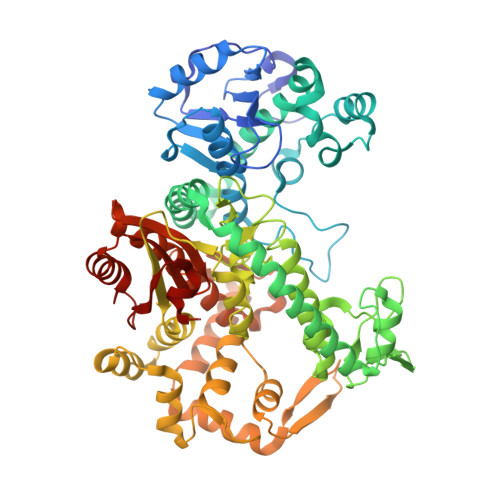
Structural dynamics and determinants of 2-aminoadenine specificity in DNA polymerase DpoZ of vibriophage phi VC8.
Czernecki, D., Hu, H., Romoli, F., Delarue, M.(2021) Nucleic Acids Res 49: 11974-11985
- PubMed: 34751404 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab955
- Primary Citation Related Structures:
7PBK - PubMed Abstract:
All genetic information in cellular life is stored in DNA copolymers composed of four basic building blocks (ATGC-DNA). In contrast, a group of bacteriophages belonging to families Siphoviridae and Podoviridae has abandoned the usage of one of them, adenine (A), replacing it with 2-aminoadenine (Z). The resulting ZTGC-DNA is more stable than its ATGC-DNA counterpart, owing to the additional hydrogen bond present in the 2-aminoadenine:thymine (Z:T) base pair, while the additional amino group also confers resistance to the host endonucleases. Recently, two classes of replicative proteins found in ZTGC-DNA-containing phages were characterized and one of them, DpoZ from DNA polymerase A (PolA) family, was shown to possess significant Z-vs-A specificity. Here, we present the crystallographic structure of the apo form of DpoZ of vibriophage ϕVC8, composed of the 3'-5' exonuclease and polymerase domains. We captured the enzyme in two conformations that involve the tip of the thumb subdomain and the exonuclease domain. We highlight insertions and mutations characteristic of ϕVC8 DpoZ and its close homologues. Through mutagenesis and functional assays we suggest that the preference of ϕVC8 DpoZ towards Z relies on a polymerase backtracking process, more efficient when the nascent base pair is A:T than when it is Z:T.
- Unit of Architecture and Dynamics of Biological Macromolecules, CNRS UMR 3528, 25-28 rue du Docteur Roux, Institut Pasteur, 75015 Paris, France.
Organizational Affiliation: