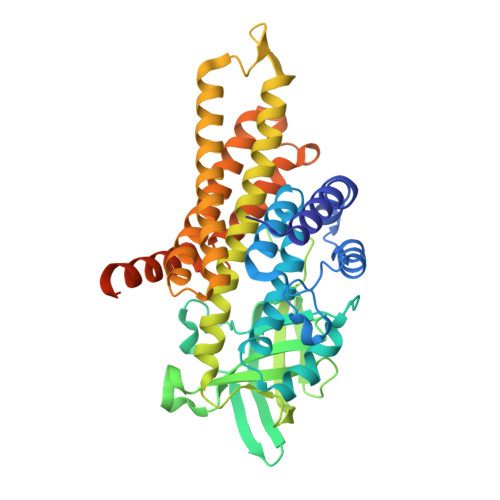
Structural Basis of Cyclic 1,3-Diene Forming Acyl-Coenzyme A Dehydrogenases.
Kung, J.W., Meier, A.K., Willistein, M., Weidenweber, S., Demmer, U., Ermler, U., Boll, M.(2021) Chembiochem 22: 3173-3177
- PubMed: 34555236 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cbic.202100421
- Primary Citation Related Structures:
7P98, 7P9A, 7P9X - PubMed Abstract:
The biologically important, FAD-containing acyl-coenzyme A (CoA) dehydrogenases (ACAD) usually catalyze the anti-1,2-elimination of a proton and a hydride of aliphatic CoA thioesters. Here, we report on the structure and function of an ACAD from anaerobic bacteria catalyzing the unprecedented 1,4-elimination at C3 and C6 of cyclohex-1-ene-1-carboxyl-CoA (Ch1CoA) to cyclohex-1,5-diene-1-carboxyl-CoA (Ch1,5CoA) and at C3 and C4 of the latter to benzoyl-CoA. Based on high-resolution Ch1CoA dehydrogenase crystal structures, the unorthodox reactivity is explained by the presence of a catalytic aspartate base (D91) at C3, and by eliminating the catalytic glutamate base at C1. Moreover, C6 of Ch1CoA and C4 of Ch1,5CoA are positioned towards FAD-N5 to favor the biologically relevant C3,C6- over the C3,C4-dehydrogenation activity. The C1,C2-dehydrogenation activity was regained by structure-inspired amino acid exchanges. The results provide the structural rationale for the extended catalytic repertoire of ACADs and offer previously unknown biocatalytic options for the synthesis of cyclic 1,3-diene building blocks.
- Faculty of Biology - Microbiology, Albert-Ludwigs-Universität Freiburg, Schänzlestrasse 1, 79104, Freiburg, Germany.
Organizational Affiliation: