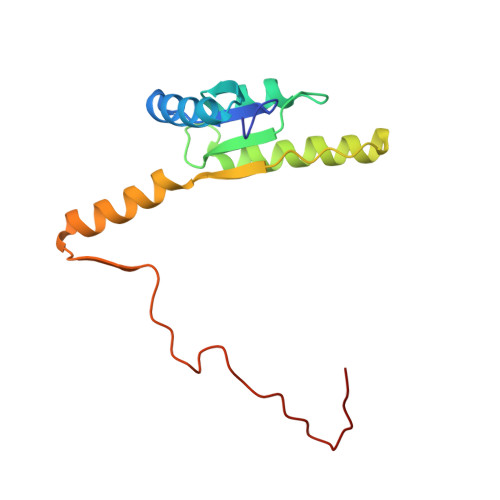
3D Domain Swapping Dimerization of the Receiver Domain of Cytokinin Receptor CRE1 From Arabidopsis thaliana and Medicago truncatula .
Tran, L.H., Urbanowicz, A., Jasinski, M., Jaskolski, M., Ruszkowski, M.(2021) Front Plant Sci 12: 756341-756341
- PubMed: 34630499
- DOI: https://doi.org/10.3389/fpls.2021.756341
- Primary Citation Related Structures:
7P8C, 7P8D, 7P8E - PubMed Abstract:
Cytokinins are phytohormones regulating many biological processes that are vital to plants. CYTOKININ RESPONSE1 (CRE1), the main cytokinin receptor, has a modular architecture composed of a cytokinin-binding CHASE (Cyclases/Histidine kinases Associated Sensory Extracellular) domain, followed by a transmembrane fragment, an intracellular histidine kinase (HK) domain, and a receiver domain (REC). Perception of cytokinin signaling involves (i) a hormone molecule binding to the CHASE domain, (ii) CRE1 autophosphorylation at a conserved His residue in the HK domain, followed by a phosphorelay to (iii) a conserved Asp residue in the REC domain, (iv) a histidine-containing phosphotransfer protein (HPt), and (v) a response regulator (RR). This work focuses on the crystal structures of the REC domain of CRE1 from the model plant Arabidopsis thaliana and from the model legume Medicago truncatula . Both REC domains form tight 3D-domain-swapped dimers. Dimerization of the REC domain agrees with the quaternary assembly of the entire CRE1 but is incompatible with a model of its complex with HPt, suggesting that a considerable conformational change should occur to enable the signal transduction. Indeed, phosphorylation of the REC domain can change the HPt-binding properties of CRE1, as shown by functional studies.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznań, Poland.
Organizational Affiliation: